6Q6F
 
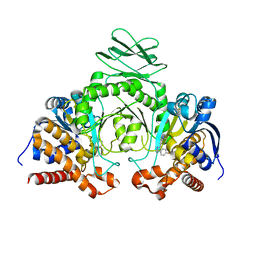 | | Crystal structure of IDH1 R132H in complex with HMS101 | | Descriptor: | (2~{R})-2-[2-[(3~{R})-3-(4-fluorophenyl)pyrrolidin-1-yl]ethyl]-1,4-dimethyl-piperazine, Isocitrate dehydrogenase [NADP] cytoplasmic | | Authors: | Chaturvedi, A, Goparaju, R, Gupta, C, Kluenemann, T, Araujo Cruz, M.M, Kloos, A, Goerlich, K, Schottmann, R, Struys, E.A, Ganser, A, Preller, M, Heuser, M. | | Deposit date: | 2018-12-10 | | Release date: | 2019-10-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | In vivo efficacy of mutant IDH1 inhibitor HMS-101 and structural resolution of distinct binding site.
Leukemia, 34, 2020
|
|
6T65
 
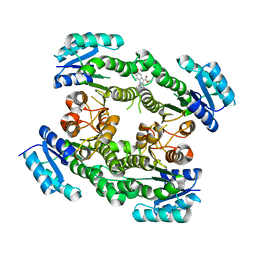 | |
6TBD
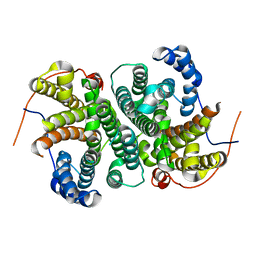 
 | |
6TJA
 
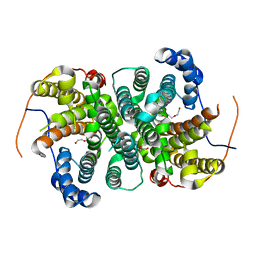 | | Crystal structure of the SVS_A2 protein (W79F,G83L mutant) from ancestral sequence reconstruction at 2.27 A resolution | | Descriptor: | DI(HYDROXYETHYL)ETHER, SVS_variant_AS1 | | Authors: | Rudraraju, R, Schnell, R, Schneider, G. | | Deposit date: | 2019-11-25 | | Release date: | 2020-12-09 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Engineering of Ancestors as a Tool to Elucidate Structure, Mechanism, and Specificity of Extant Terpene Cyclase.
J.Am.Chem.Soc., 143, 2021
|
|
6TIV
 
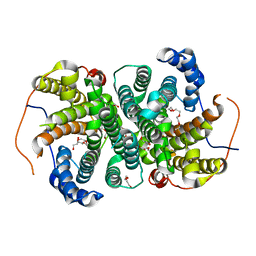 | | Crystal structure of the SVS_A2 protein (205-DREMH-209 /205-AQDLE-209 mutant) from ancestral sequence reconstruction at 2.38 A resolution | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, SVS variant AT2, ... | | Authors: | Rudraraju, R, Schnell, R, Schneider, G. | | Deposit date: | 2019-11-22 | | Release date: | 2020-12-02 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Engineering of Ancestors as a Tool to Elucidate Structure, Mechanism, and Specificity of Extant Terpene Cyclase.
J.Am.Chem.Soc., 143, 2021
|
|
6TJZ
 
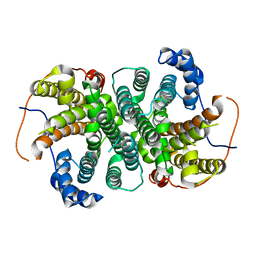 | |
6THU
 
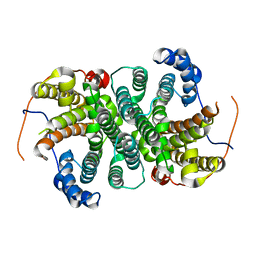 | |
8WSV
 
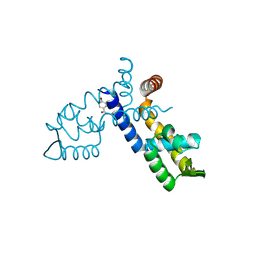 | |
8XB7
 
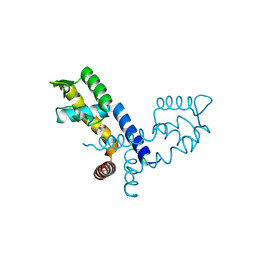 | |
5D7E
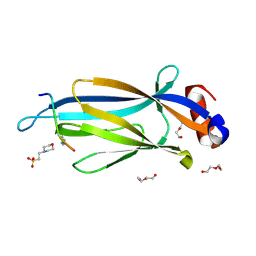 
 | | Crystal structure of Taf14 YEATS domain in complex with H3K9ac | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, H3K9ac, ... | | Authors: | Andrews, F.H, Shanle, E.K, Strahl, B.D, Kutateladze, T.G. | | Deposit date: | 2015-08-13 | | Release date: | 2015-09-23 | | Last modified: | 2019-12-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Association of Taf14 with acetylated histone H3 directs gene transcription and the DNA damage response.
Genes Dev., 29, 2015
|
|
