3EJ6
 
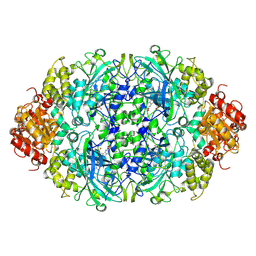 | | Neurospora Crassa Catalase-3 Crystal Structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Catalase-3, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Diaz, A, Valdes, V.-J, Rudino-Pinera, E, Horjales, E, Hansberg, W. | | Deposit date: | 2008-09-17 | | Release date: | 2009-01-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure-Function Relationships in Fungal Large-Subunit Catalases
J.Mol.Biol., 386, 2008
|
|
1SY7
 
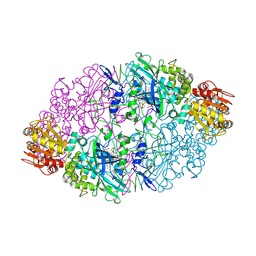 | | Crystal structure of the catalase-1 from Neurospora crassa, native structure at 1.75A resolution. | | Descriptor: | CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, Catalase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Diaz, A, Horjales, E, Rudino-Pinera, E, Arreola, R, Hansberg, W. | | Deposit date: | 2004-04-01 | | Release date: | 2004-10-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Unusual Cys-Tyr covalent bond in a large catalase
J.Mol.Biol., 342, 2004
|
|
5WHS
 
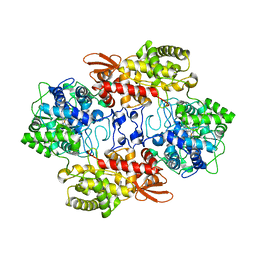 | | Crystal structure of the catalase-peroxidase from Neurospora crassa at 2.6 A | | Descriptor: | Catalase-peroxidase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Diaz-Vilchis, A, Vega-Garcia, V, Rudino-Pinera, E, Hansberg, W. | | Deposit date: | 2017-07-18 | | Release date: | 2018-01-17 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure, kinetics, molecular and redox properties of a cytosolic and developmentally regulated fungal catalase-peroxidase.
Arch. Biochem. Biophys., 640, 2018
|
|
5WHQ
 
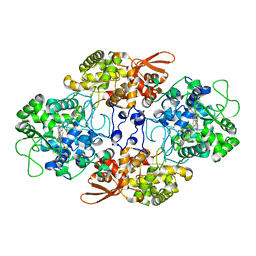 | | Crystal structure of the catalase-peroxidase from Neurospora crassa at 2.9 A | | Descriptor: | Catalase-peroxidase, POTASSIUM ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Diaz-Vilchis, A, Vega-Garcia, V, Rudino-Pinera, E, Hansberg, W. | | Deposit date: | 2017-07-18 | | Release date: | 2018-01-17 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure, kinetics, molecular and redox properties of a cytosolic and developmentally regulated fungal catalase-peroxidase.
Arch. Biochem. Biophys., 640, 2018
|
|
4BIM
 
 | |
3ZJ4
 
 | |
3ZJ5
 
 | | NEUROSPORA CRASSA CATALASE-3 EXPRESSED IN E. COLI, ORTHORHOMBIC FORM. | | Descriptor: | 1,2-ETHANEDIOL, 2-(2-ETHOXYETHOXY)ETHANOL, CATALASE-3, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E. | | Deposit date: | 2013-01-17 | | Release date: | 2013-07-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Conformational Stability and Crystal Packing: Polymorphism in Neurospora Crassa Cat-3
Acta Crystallogr.,Sect.F, 69, 2013
|
|
4AJ9
 
 | | Catalase 3 from Neurospora crassa | | Descriptor: | ACETATE ION, CATALASE-3, PENTAETHYLENE GLYCOL, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E. | | Deposit date: | 2012-02-16 | | Release date: | 2013-03-06 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | X-ray driven reduction of Cpd I of Catalase-3 from N. crassa reveals differential sensitivity of active sites and formation of ferrous state.
Arch.Biochem.Biophys., 666, 2019
|
|
6NSW
 
 | | X-ray reduced Catalase 3 From N.Crassa in Cpd I state (0.135 MGy) | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Catalase-3, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E, Stojanoff, V. | | Deposit date: | 2019-01-25 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.099 Å) | | Cite: | X-ray driven reduction of Cpd I of Catalase-3 from N. crassa reveals differential sensitivity of active sites and formation of ferrous state.
Arch.Biochem.Biophys., 666, 2019
|
|
6NSZ
 
 | | X-ray reduced Catalase 3 from N.Crassa (0.526 MGy) | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Catalase-3, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E, Stojanoff, V. | | Deposit date: | 2019-01-27 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray driven reduction of Cpd I of Catalase-3 from N. crassa reveals differential sensitivity of active sites and formation of ferrous state.
Arch.Biochem.Biophys., 666, 2019
|
|
6NT1
 
 | | Catalase 3 from N.Crassa in ferrous state (2.89 MGy) | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Catalase-3, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E, Stojanoff, V. | | Deposit date: | 2019-01-27 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray driven reduction of Cpd I of Catalase-3 from N. crassa reveals differential sensitivity of active sites and formation of ferrous state.
Arch.Biochem.Biophys., 666, 2019
|
|
6NSY
 
 | | X-ray reduced Catalase 3 From N.Crassa in Cpd I state (0.263 MGy) | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Catalase-3, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E, Stojanoff, V. | | Deposit date: | 2019-01-27 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray driven reduction of Cpd I of Catalase-3 from N. crassa reveals differential sensitivity of active sites and formation of ferrous state.
Arch.Biochem.Biophys., 666, 2019
|
|
6NT0
 
 | | Catalase 3 from N.Crassa in ferrous state, X-ray reduced (1.315 MGy) | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Catalase-3, ... | | Authors: | Zarate-Romero, A, Rudino-Pinera, E, Stojanoff, V. | | Deposit date: | 2019-01-27 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray driven reduction of Cpd I of Catalase-3 from N. crassa reveals differential sensitivity of active sites and formation of ferrous state.
Arch.Biochem.Biophys., 666, 2019
|
|
