1F39
 
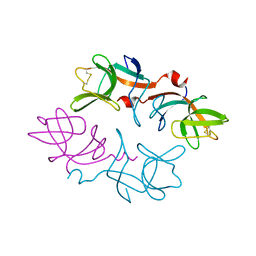 | | CRYSTAL STRUCTURE OF THE LAMBDA REPRESSOR C-TERMINAL DOMAIN | | Descriptor: | REPRESSOR PROTEIN CI | | Authors: | Bell, C.E, Frescura, P, Hochschild, A, Lewis, M. | | Deposit date: | 2000-06-01 | | Release date: | 2000-07-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the lambda repressor C-terminal domain provides a model for cooperative operator binding.
Cell(Cambridge,Mass.), 101, 2000
|
|
1SGK
 
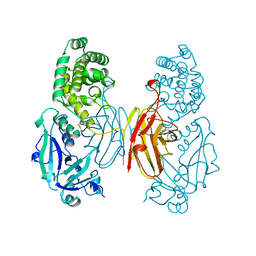 | | NUCLEOTIDE-FREE DIPHTHERIA TOXIN | | Descriptor: | DIPHTHERIA TOXIN (DIMERIC) | | Authors: | Bell, C.E, Eisenberg, D. | | Deposit date: | 1996-09-12 | | Release date: | 1996-12-23 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of nucleotide-free diphtheria toxin.
Biochemistry, 36, 1997
|
|
1TOX
 
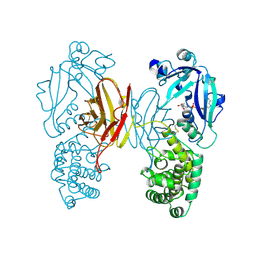 | | DIPHTHERIA TOXIN DIMER COMPLEXED WITH NAD | | Descriptor: | DIPHTHERIA TOXIN (DIMERIC), NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Bell, C.E, Eisenberg, D. | | Deposit date: | 1995-10-06 | | Release date: | 1996-06-10 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of diphtheria toxin bound to nicotinamide adenine dinucleotide.
Biochemistry, 35, 1996
|
|
1EFA
 
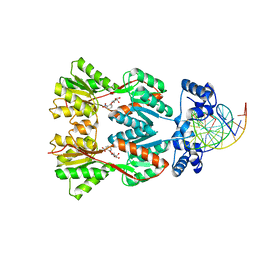 | |
1JYE
 
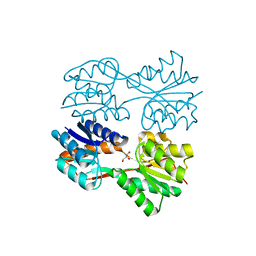 | | Structure of a Dimeric Lac Repressor with C-terminal Deletion and K84L Substitution | | Descriptor: | GLYCEROL, Lactose Operon Repressor | | Authors: | Bell, C.E, Barry, J, Matthews, K.S, Lewis, M. | | Deposit date: | 2001-09-12 | | Release date: | 2001-10-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of a variant of lac repressor with increased thermostability and decreased affinity for operator.
J.Mol.Biol., 313, 2001
|
|
1KCA
 
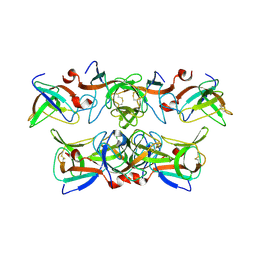 | |
1JWL
 
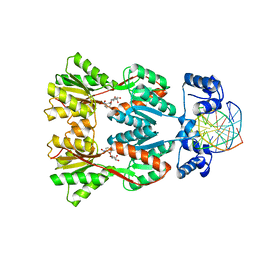 | | Structure of the Dimeric lac Repressor/Operator O1/ONPF Complex | | Descriptor: | 2-nitrophenyl beta-D-fucopyranoside, 5'-D(*AP*GP*AP*AP*T*TP*GP*TP*GP*AP*GP*CP*GP*GP*AP*TP*AP*AP*CP*AP*AP*TP*T)-3', 5'-D(*TP*AP*AP*TP*TP*GP*TP*TP*AP*TP*CP*CP*GP*CP*TP*CP*AP*CP*AP*AP*TP*TP*C)-3', ... | | Authors: | Bell, C.E, Lewis, M. | | Deposit date: | 2001-09-04 | | Release date: | 2001-10-05 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Crystallographic analysis of Lac repressor bound to natural operator O1.
J.Mol.Biol., 312, 2001
|
|
1JYF
 
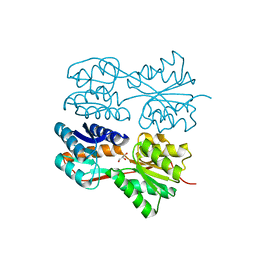 | | Structure of the Dimeric Lac Repressor with an 11-residue C-terminal Deletion. | | Descriptor: | GLYCEROL, Lactose Operon Repressor | | Authors: | Bell, C.E, Barry, J, Matthews, K.S, Lewis, M. | | Deposit date: | 2001-09-12 | | Release date: | 2001-10-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of a variant of lac repressor with increased thermostability and decreased affinity for operator.
J.Mol.Biol., 313, 2001
|
|
4JRP
 
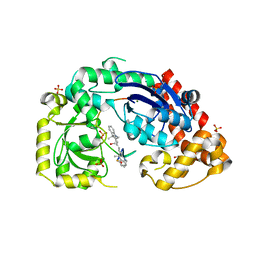 | | Structure of E. coli Exonuclease I in complex with a 5cy-dT13 oligonucleotide | | Descriptor: | 5cy-dT13, Exodeoxyribonuclease I, GLYCEROL, ... | | Authors: | Bell, C.E. | | Deposit date: | 2013-03-21 | | Release date: | 2013-05-08 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structures of Escherichia coli exonuclease I in complex with single-stranded DNA provide insights into the mechanism of processive digestion.
Nucleic Acids Res., 41, 2013
|
|
4JRQ
 
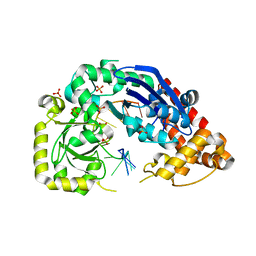 | |
4JS5
 
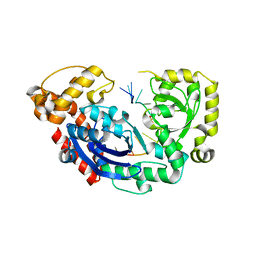 | |
4JS4
 
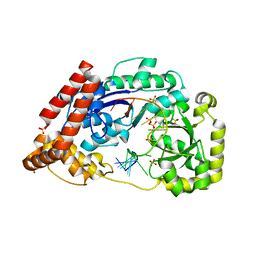 | |
8TG8
 
 | |
8TG7
 
 | |
8TGC
 
 | |
8TFU
 
 | |
3H4R
 
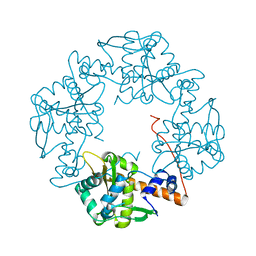 | | Crystal structure of E. coli RecE exonuclease | | Descriptor: | Exodeoxyribonuclease 8 | | Authors: | Bell, C.E, Zhang, J. | | Deposit date: | 2009-04-20 | | Release date: | 2009-05-26 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of E. coli RecE protein reveals a toroidal tetramer for processing double-stranded DNA breaks.
Structure, 17, 2009
|
|
1XMS
 
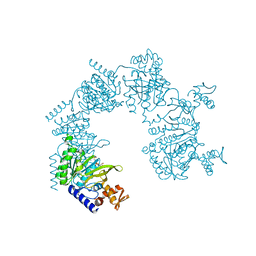 | | E. coli RecA in complex with MnAMP-PNP | | Descriptor: | MANGANESE (II) ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, RecA protein | | Authors: | Bell, C.E, Xing, X. | | Deposit date: | 2004-10-04 | | Release date: | 2005-01-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structures of Escherichia coli RecA in Complex with MgADP and MnAMP-PNP(,).
Biochemistry, 43, 2004
|
|
1XMV
 
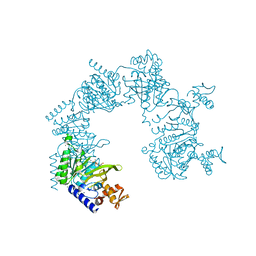 | | E. coli RecA in complex with MgADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, RecA protein | | Authors: | Bell, C.E, Xing, X. | | Deposit date: | 2004-10-04 | | Release date: | 2005-01-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structures of Escherichia coli RecA in Complex with MgADP and MnAMP-PNP(,).
Biochemistry, 43, 2004
|
|
1XP8
 
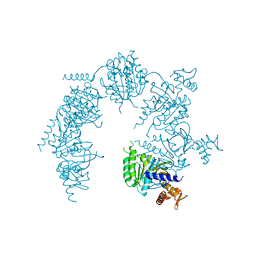 | | Deinococcus radiodurans RecA in complex with ATP-gamma-S | | Descriptor: | PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, RecA protein | | Authors: | Bell, C.E, Rajan, R. | | Deposit date: | 2004-10-08 | | Release date: | 2004-12-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of RecA from Deinococcus radiodurans: insights into the structural basis of extreme radioresistance.
J.Mol.Biol., 344, 2004
|
|
3SM4
 
 | |
3SLP
 
 | |
6M9K
 
 | |
8GLR
 
 | | FrlB - Deglycase for 6-phosphofructose lysine | | Descriptor: | GLYCEROL, SIS domain-containing protein | | Authors: | Bell, C.E, Gopalan, V, Kovvali, S. | | Deposit date: | 2023-03-22 | | Release date: | 2023-07-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Insights into the catalytic mechanism of a bacterial deglycase essential for utilization of fructose-lysine.
Protein Sci., 32, 2023
|
|
7UBB
 
 | |
