7TJ3
 
 | |
6UDF
 
 | |
6UJ5
 
 | |
6UJK
 
 | |
6ULO
 
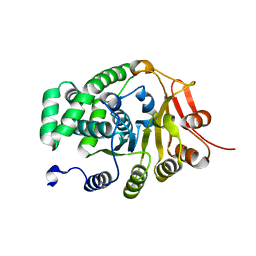 | |
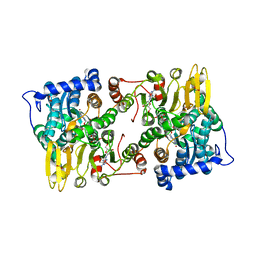
6UK3
 
 | |
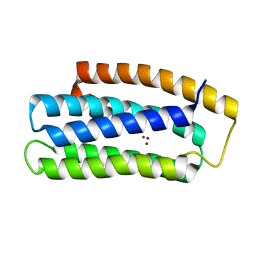
6TYJ
 
 | |
6UCZ
 
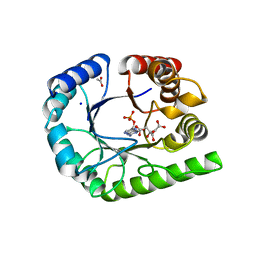 | |
6UDE
 | |
6UH2
 
 | |
6ULD
 
 | |
6UDG
 
 | |
7ULH
 
 | |
7ULZ
 
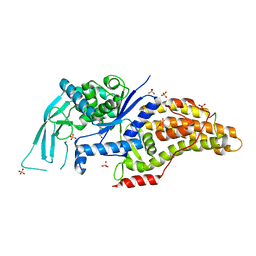 | |
7V0H
 | |
4DZ3
 
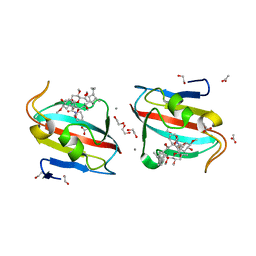 | |
3T7C
 
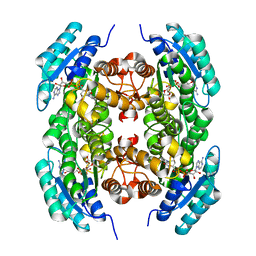 | |
3SX2
 
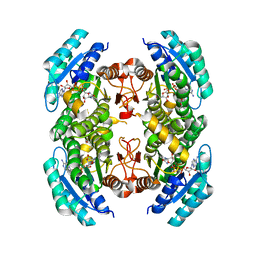 | |
3TK1
 
 | |
3U0B
 
 | | Crystal structure of an oxidoreductase from Mycobacterium smegmatis | | Descriptor: | Oxidoreductase, short chain dehydrogenase/reductase family protein, SODIUM ION | | Authors: | Arakaki, T.L, Staker, B.L, Clifton, M.C, Abendroth, J, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-09-28 | | Release date: | 2011-10-05 | | Last modified: | 2024-07-17 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and functional characterization of FabG4 from Mycolicibacterium smegmatis.
Acta Crystallogr.,Sect.F, 80, 2024
|
|
3TSC
 
 | |
8DV0
 
 | |
8DTC
 
 | |
8DU0
 
 | |
8DT6
 
 | |
