7KAY
 
 | |
7KAX
 
 | |
7KAZ
 
 | |
7KAV
 
 | |
7KAW
 
 | |
8VDA
 
 | |
8VD9
 
 | |
8VDB
 
 | |
9AXE
 
 | |
8UM2
 | |
8UM1
 
 | |
8UR6
 
 | |
8UR3
 
 | |
7UQ1
 
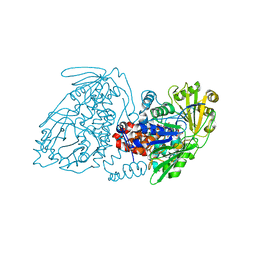
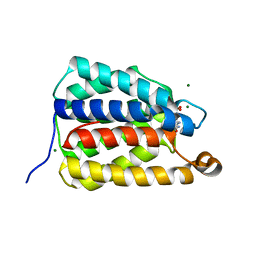 | | The X-ray crystal structure of the N-terminal domain of Staphylococcus aureus Fatty Acid Kinase A (FakA, residues 1-208) in complex with AMP and a single Mg ion at the dinuclear binding site | | Descriptor: | ADENOSINE MONOPHOSPHATE, Fatty Acid Kinase A, GLYCEROL, ... | | Authors: | Cuypers, M.G, Subramanian, C, Seetharaman, J, Rock, C.O, White, S.W. | | Deposit date: | 2022-04-18 | | Release date: | 2022-05-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Domain architecture and catalysis of the Staphylococcus aureus fatty acid kinase.
J.Biol.Chem., 298, 2022
|
|
8UR8
 
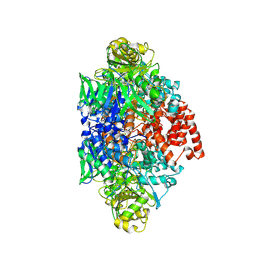 | |
8TQE
 
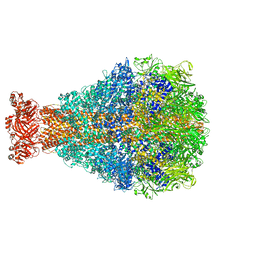 | | XptA2 wild type | | Descriptor: | XptA2 | | Authors: | Martin, C.L, Binshtein, E.M, Aller, S.G. | | Deposit date: | 2023-08-07 | | Release date: | 2023-09-27 | | Last modified: | 2023-10-11 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structures of the Insecticidal Toxin Complex Subunit XptA2 Highlight Roles for Flexible Domains.
Int J Mol Sci, 24, 2023
|
|
8TV0
 
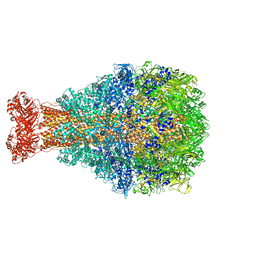 | | XptA2 wild type | | Descriptor: | XptA2 | | Authors: | Martin, C.L, Aller, S.G. | | Deposit date: | 2023-08-17 | | Release date: | 2023-09-27 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structures of the Insecticidal Toxin Complex Subunit XptA2 Highlight Roles for Flexible Domains.
Int J Mol Sci, 24, 2023
|
|
