5CCO
 
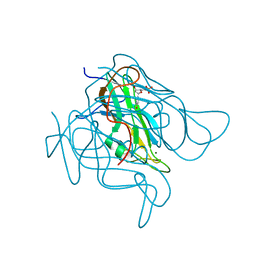 | | Staphylococcus bacteriophage 80alpha dUTPase with dUMP. | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, DUTPase, MAGNESIUM ION | | Authors: | Maiques, E, Quiles-Puchalt, N, Donderis, J, Ciges, J.R, Alite, C, Bowring, J, Penades, J.R, Marina, A. | | Deposit date: | 2015-07-02 | | Release date: | 2016-05-11 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Another look at the mechanism involving trimeric dUTPases in Staphylococcus aureus pathogenicity island induction involves novel players in the party.
Nucleic Acids Res., 44, 2016
|
|
5CCT
 
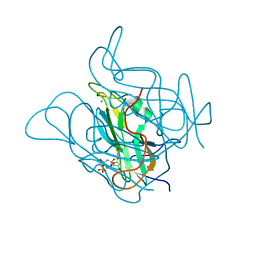 | | Staphylococcus bacteriophage 80alpha dUTPase G164S mutant with dUpNHpp. | | Descriptor: | 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, DUTPase, MAGNESIUM ION | | Authors: | Maiques, E, Quiles-Puchalt, N, Donderis, J, Ciges, J.R, Alite, C, Bowring, J, Penades, J.R, Marina, A. | | Deposit date: | 2015-07-02 | | Release date: | 2016-05-11 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Another look at the mechanism involving trimeric dUTPases in Staphylococcus aureus pathogenicity island induction involves novel players in the party.
Nucleic Acids Res., 44, 2016
|
|
7Q0N
 
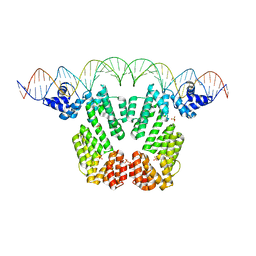 | | Arbitrium receptor from Katmira phage | | Descriptor: | Arbitrium receptor, DNA (45-MER), SULFATE ION | | Authors: | Gallego del Sol, F, Marina, A. | | Deposit date: | 2021-10-15 | | Release date: | 2022-06-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Insights into the mechanism of action of the arbitrium communication system in SPbeta phages.
Nat Commun, 13, 2022
|
|
6S7I
 
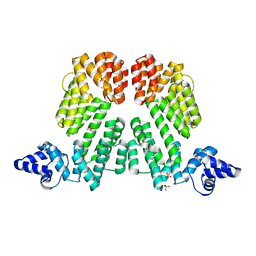 | |
6S7L
 
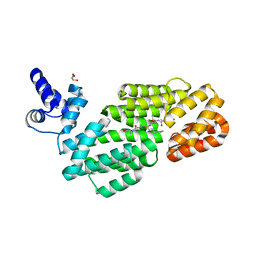 | |
8ANT
 
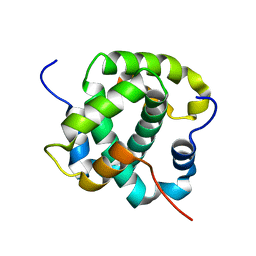 | |
8ANU
 
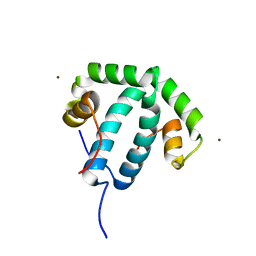 | | Crystal structure of protein phi3T-93 | | Descriptor: | NICKEL (II) ION, YopN. Phi3T_93 | | Authors: | Zamora-Caballero, S, Marina, A. | | Deposit date: | 2022-08-05 | | Release date: | 2023-10-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.310147 Å) | | Cite: | Antagonistic interactions between phage and host factors control arbitrium lysis-lysogeny decision.
Nat Microbiol, 9, 2024
|
|
8ANV
 
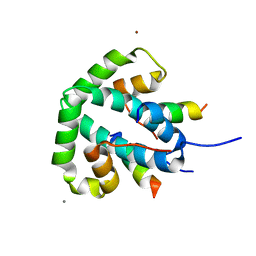 | |
8BJ6
 
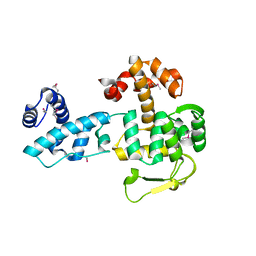 | | Crystal structure of YopR | | Descriptor: | SPbeta prophage-derived uncharacterized protein YopR | | Authors: | Gallego del Sol, F, Marina, A. | | Deposit date: | 2022-11-03 | | Release date: | 2023-11-15 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Characterization of a unique repression system present in arbitrium phages of the SPbeta family.
Cell Host Microbe, 31, 2023
|
|
8BJV
 
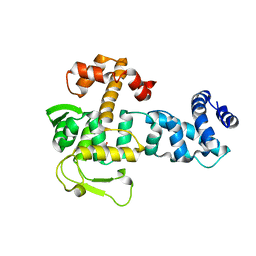 | | Crystal structure of YopR | | Descriptor: | GLYCEROL, SPbeta prophage-derived uncharacterized protein YopR | | Authors: | Gallego del Sol, F, Marina, A. | | Deposit date: | 2022-11-08 | | Release date: | 2023-11-22 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Characterization of a unique repression system present in arbitrium phages of the SPbeta family.
Cell Host Microbe, 31, 2023
|
|
8BPZ
 
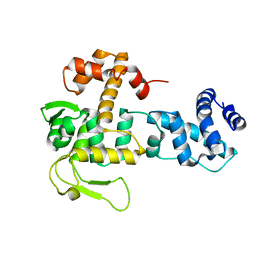 | | Crystal structure of YopR | | Descriptor: | SPbeta prophage-derived uncharacterized protein YopR | | Authors: | Gallego del Sol, F, Marina, A. | | Deposit date: | 2022-11-18 | | Release date: | 2023-11-29 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Characterization of a unique repression system present in arbitrium phages of the SPbeta family.
Cell Host Microbe, 31, 2023
|
|
8C8E
 
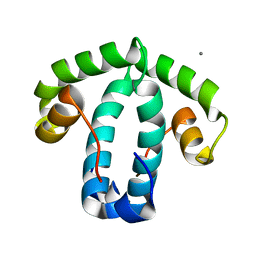 | |
