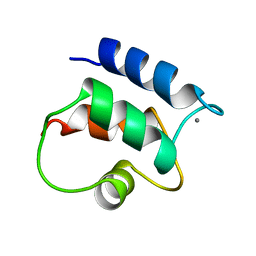
1B1G
 
 | |
1BKF
 
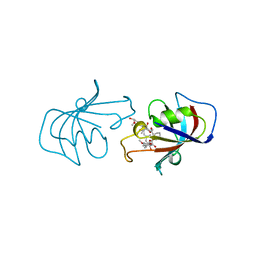 | | FK506 BINDING PROTEIN FKBP MUTANT R42K/H87V COMPLEX WITH IMMUNOSUPPRESSANT FK506 | | Descriptor: | 8-DEETHYL-8-[BUT-3-ENYL]-ASCOMYCIN, FK506 BINDING PROTEIN | | Authors: | Itoh, S, Decenzo, M.T, Livingston, D.J, Pearlman, D.A, Navia, M.A. | | Deposit date: | 1995-10-18 | | Release date: | 1996-08-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Conformation of Fk506 in X-Ray Structures of its Complexes with Human Recombinant Fkbp12 Mutants
Bioorg.Med.Chem.Lett., 5, 1995
|
|
3FC1
 
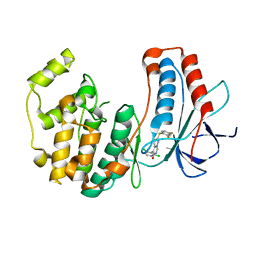 | | Crystal structure of p38 kinase bound to pyrimido-pyridazinone inhibitor | | Descriptor: | 5-(2,6-dichlorophenyl)-2-[(2,4-difluorophenyl)sulfanyl]-6H-pyrimido[1,6-b]pyridazin-6-one, CHLORIDE ION, Mitogen-activated protein kinase 14 | | Authors: | Jacobs, M.D, Bellon, S. | | Deposit date: | 2008-11-20 | | Release date: | 2008-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Evaluating the molecular mechanics poisson-boltzmann surface area free energy method using a congeneric series of ligands to p38 MAP kinase.
J.Med.Chem., 48, 2005
|
|
1D42
 
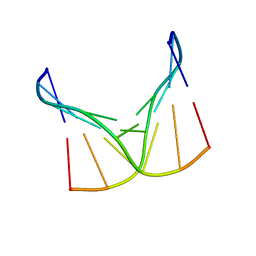 | |
