2MYI
 
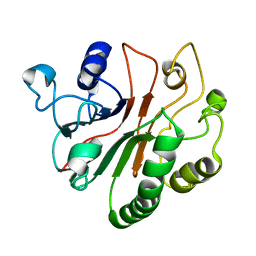 | | Solution Structure of Crc from P. syringae Lz4W | | Descriptor: | Exodeoxyribonuclease III | | Authors: | Deshmukh, M.V, Sharma, R. | | Deposit date: | 2015-01-25 | | Release date: | 2016-01-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Crc of Pseudomonas syringae Lz4W utilizes the dominant RNA binding site for mutually exclusive interactions with Hfq:mRNA and CrcY/Z RNA
J.Magn.Reson., 10-11, 2022
|
|
8IGD
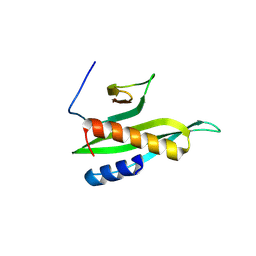 
 | |
8KFP
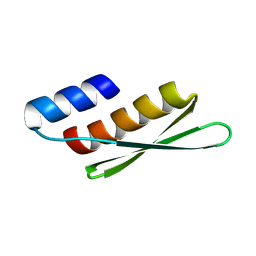 
 | |
2JVB
 
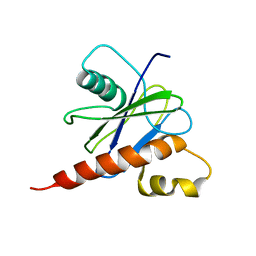 | |
2LTS
 
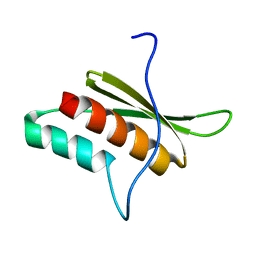 | | Solution structure of RDE-4(150-235) | | Descriptor: | Protein RDE-4 | | Authors: | Deshmukh, M, Chiliveri, S. | | Deposit date: | 2012-05-31 | | Release date: | 2013-12-04 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure of RDE-4 dsRBDs and mutational studies provide insights into dsRNA recognition in the Caenorhabditis elegans RNAi pathway.
Biochem.J., 458, 2014
|
|
2LTR
 
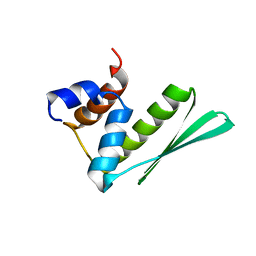 | | Solution structure of RDE-4(32-136) | | Descriptor: | Protein RDE-4 | | Authors: | Deshmukh, M, Chiliveri, S. | | Deposit date: | 2012-05-31 | | Release date: | 2013-12-04 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure of RDE-4 dsRBDs and mutational studies provide insights into dsRNA recognition in the Caenorhabditis elegans RNAi pathway.
Biochem.J., 458, 2014
|
|
4NBJ
 
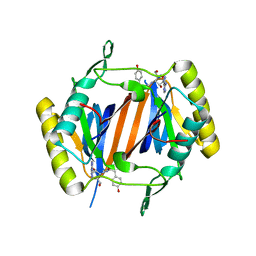 | | D-aminoacyl-tRNA deacylase (DTD) from Plasmodium falciparum in complex with D-tyrosyl-3'-aminoadenosine at 2.20 Angstrom resolution | | Descriptor: | 3'-deoxy-3'-(D-tyrosylamino)adenosine, D-tyrosyl-tRNA(Tyr) deacylase | | Authors: | Ahmad, S, Routh, S.B, Kamarthapu, V, Sankaranarayanan, R. | | Deposit date: | 2013-10-23 | | Release date: | 2013-12-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanism of chiral proofreading during translation of the genetic code.
Elife, 2, 2013
|
|
4NBI
 
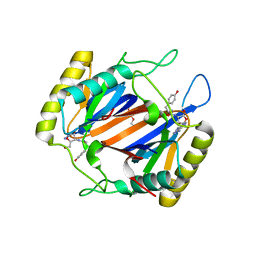 | | D-aminoacyl-tRNA deacylase (DTD) from Plasmodium falciparum in complex with D-tyrosyl-3'-aminoadenosine at 1.86 Angstrom resolution | | Descriptor: | 3'-deoxy-3'-(D-tyrosylamino)adenosine, D-tyrosyl-tRNA(Tyr) deacylase, TRIETHYLENE GLYCOL | | Authors: | Ahmad, S, Routh, S.B, Kamarthapu, V, Sankaranarayanan, R. | | Deposit date: | 2013-10-23 | | Release date: | 2013-12-18 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Mechanism of chiral proofreading during translation of the genetic code.
Elife, 2, 2013
|
|
2N3G
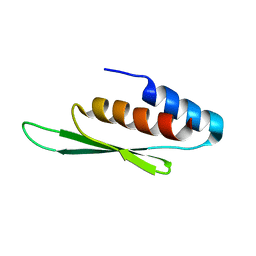 
 | |
2N3H
 
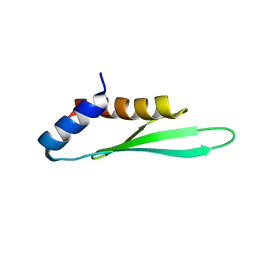 | |
2N3F
 
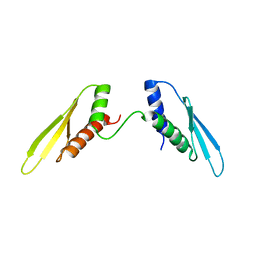 | |
3PD4
 
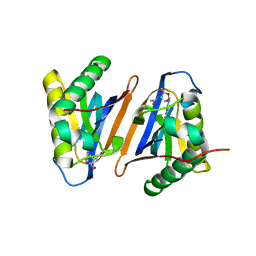 | |
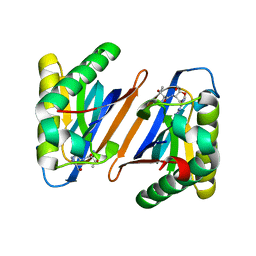
3PD3
 
 | |
3PD2
 
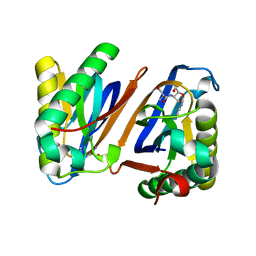 | | Crystal structure of the editing domain of threonyl-tRNA synthetase from Pyrococcus abyssi in complex with seryl-3'-aminoadenosine | | Descriptor: | SERINE-3'-AMINOADENOSINE, Threonyl-tRNA synthetase | | Authors: | Hussain, T, Kamarthapu, V, Kruparani, S.P, Sankaranarayanan, R. | | Deposit date: | 2010-10-22 | | Release date: | 2010-12-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Mechanistic insights into cognate substrate discrimination during proofreading in translation
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3PD5
 
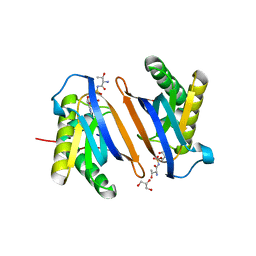 | | Crystal structure of the editing domain of threonyl-tRNA synthetase from Pyrococcus abyssi in complex with an analog of threonyl-adenylate | | Descriptor: | 5'-O-(N-(L-THREONYL)-SULFAMOYL)ADENOSINE, GLYCEROL, Threonyl-tRNA synthetase | | Authors: | Hussain, T, Kamarthapu, V, Kruparani, S.P, Sankaranarayanan, R. | | Deposit date: | 2010-10-22 | | Release date: | 2010-12-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Mechanistic insights into cognate substrate discrimination during proofreading in translation
Proc.Natl.Acad.Sci.USA, 2010
|
|
3QMM
 
 | |
