5NE6
 
 | |
6QE2
 
 | | Crystal structure of Paleococcus ferrophilus monoacylglycerol lipase. | | Descriptor: | GLYCEROL, LAURYL DIMETHYLAMINE-N-OXIDE, Monoacylglycerol lipase | | Authors: | Labar, G, Demarez, M, Brandt, N, Wouters, J, Leherte, F. | | Deposit date: | 2019-01-04 | | Release date: | 2020-02-05 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.7461 Å) | | Cite: | Structure and Dynamics of an Archeal Monoglyceride Lipase from Palaeococcus ferrophilus as Revealed by Crystallography and In Silico Analysis.
Biomolecules, 11, 2021
|
|
5L6Z
 
 | |
6NW5
 
 | |
4P6Y
 | |
5NE9
 
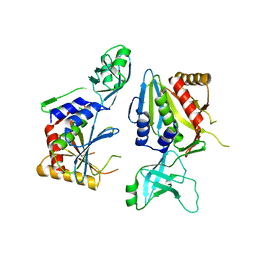 | |
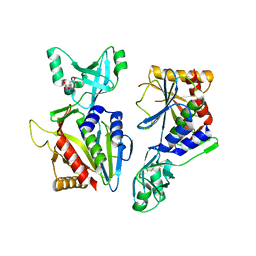
5NE7
 
 | |
5NE8
 
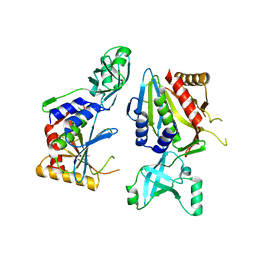 | |
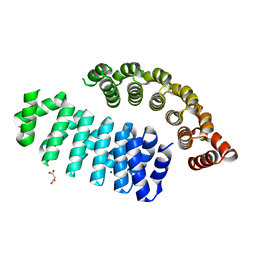
6HWP
 
 | |
6FSQ
 
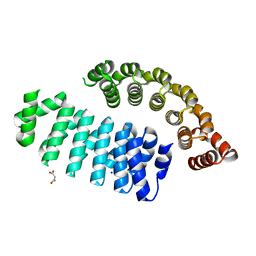 | |
6FT5
 
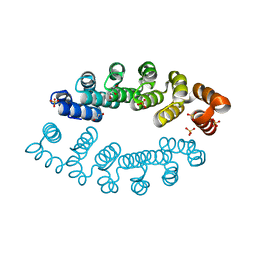 | | Structure of A3_A3, an artificial bi-domain protein based on two identical alphaRep A3 domains | | Descriptor: | GLYCEROL, SULFATE ION, alphaRep A3_A3 | | Authors: | Li de la Sierra-Gallay, I, Leger, C, Di Meo, T. | | Deposit date: | 2018-02-20 | | Release date: | 2018-08-08 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Ligand-induced conformational switch in an artificial bidomain protein scaffold.
Sci Rep, 9, 2019
|
|
