9I65
 
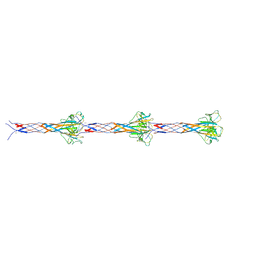 | | Recombinant F-ENA-2 fibers | | Descriptor: | DUF4183 domain-containing protein | | Authors: | Sleutel, M, Remaut, H. | | Deposit date: | 2025-01-29 | | Release date: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Cryo-EM analysis of the Bacillus thuringiensis extrasporal matrix identifies F-ENA as a widespread family of endospore appendages across the Firmicutes phylum.
Biorxiv, 2025
|
|
9I0Y
 
 | | Recombinant Ena2A fibers | | Descriptor: | DUF3992 domain-containing protein | | Authors: | Sleutel, M, Remaut, H. | | Deposit date: | 2025-01-15 | | Release date: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Cryo-EM analysis of the Bacillus thuringiensis extrasporal matrix identifies F-ENA as a widespread family of endospore appendages across the Firmicutes phylum.
Biorxiv, 2025
|
|
9H38
 
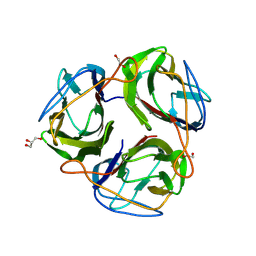 | |
9H3D
 
 | |
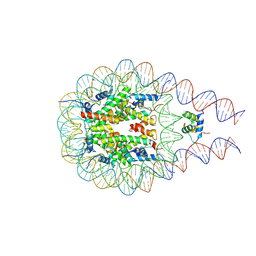
7PFV
 
 | |
7PFW
 | |
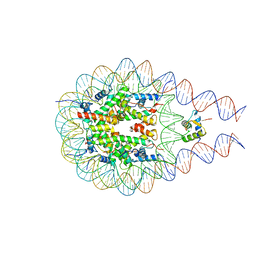
7PFX
 
 | |
7PFC
 
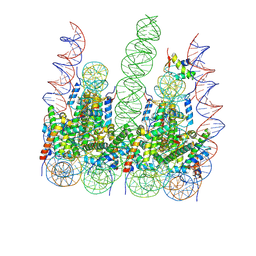 | |
7PEW
 
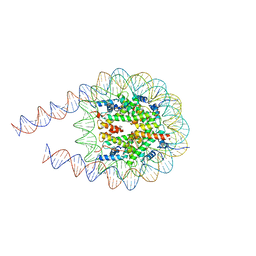 | |
7PEY
 
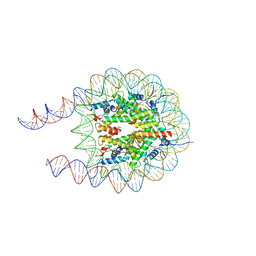 | |
7PET
 
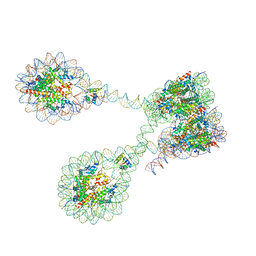 | | The 4x177 nucleosome array containing H1 | | Descriptor: | DNA (702-MER), Histone H1.4, Histone H2A type 1-B/E, ... | | Authors: | Dombrowski, M, Cramer, P. | | Deposit date: | 2021-08-11 | | Release date: | 2022-08-03 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (9.5 Å) | | Cite: | Histone H1 binding to nucleosome arrays depends on linker DNA length and trajectory.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7PEX
 
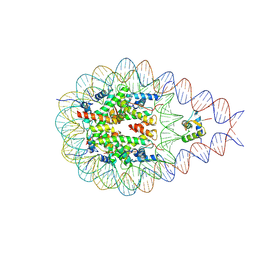 | |
7PEZ
 
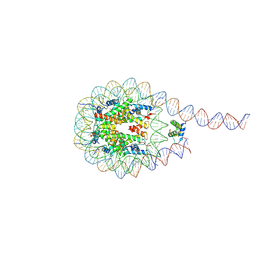 | |
7PFU
 
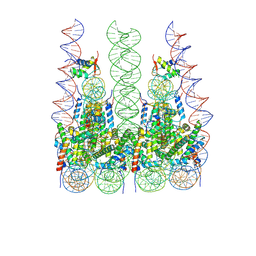 | |
7PF3
 
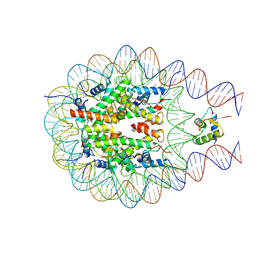 | |
7PF5
 
 | |
7PFA
 
 | |
7PF4
 
 | |
7PF6
 
 | |
7PEV
 
 | |
7PFT
 
 | |
7PF2
 
 | |
7PFF
 
 | |
7PEU
 
 | |
7PFD
 
 | |
