3CGG
 
 | |
3CSW
 
 | |
3D4O
 
 | |
3B9T
 
 | |
4NTL
 
 | |
3CLM
 
 | |
3CH0
 
 | |
3CJX
 
 | |
3CLO
 
 | |
3CSV
 
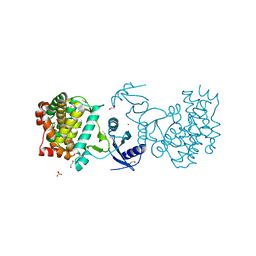 | |
4POI
 
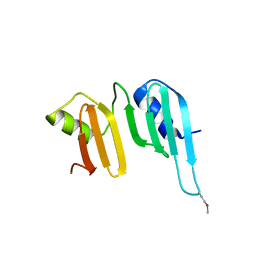 | |
3CK1
 
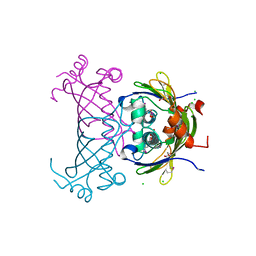 | |
3CMB
 
 | | Crystal structure of acetoacetate decarboxylase (YP_001047042.1) from Methanoculleus marisnigri JR1 at 1.60 A resolution | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, Acetoacetate decarboxylase, ... | | Authors: | Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-03-21 | | Release date: | 2008-04-01 | | Last modified: | 2023-02-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of acetoacetate decarboxylase (YP_001047042.1) from Methanoculleus marisnigri JR1 at 1.60 A resolution
To be published
|
|
3CE9
 
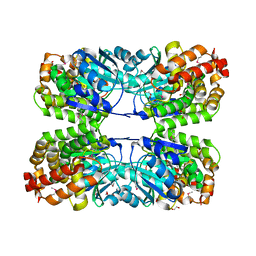 | |
4LGC
 
 | |
4LER
 
 | |
3BBY
 
 | |
4L8J
 
 | |
3BDD
 
 | |
3BEM
 | |
3B7C
 
 | |
4ME8
 
 | |
3BF4
 
 | |
4NZJ
 
 | |
3C26
 
 | |
