4A5X
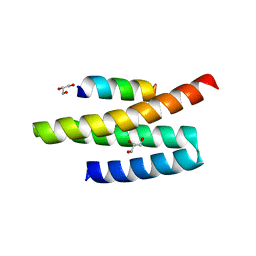 
 | | Structures of MITD1 | | Descriptor: | 2,5,8,11,14,17-HEXAOXANONADECAN-19-OL, CHARGED MULTIVESICULAR BODY PROTEIN 1A, GLYCEROL, ... | | Authors: | Hadders, M.A, Agromayor, M, Caballe, A, Obita, T, Perisic, O, Williams, R.L, Martin-Serrano, J. | | Deposit date: | 2011-10-28 | | Release date: | 2012-11-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Escrt-III Binding Protein Mitd1 is Involved in Cytokinesis and Has an Unanticipated Pld Fold that Binds Membranes.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4A5Z
 
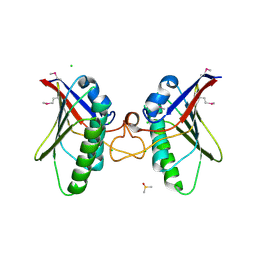 | | Structures of MITD1 | | Descriptor: | CHLORIDE ION, DIMETHYL SULFOXIDE, MIT DOMAIN-CONTAINING PROTEIN 1 | | Authors: | Hadders, M.A, Agromayor, M, Caballe, A, Obita, T, Perisic, O, Williams, R.L, Martin-Serrano, J. | | Deposit date: | 2011-10-30 | | Release date: | 2012-11-14 | | Last modified: | 2013-11-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Escrt-III Binding Protein Mitd1 is Involved in Cytokinesis and Has an Unanticipated Pld Fold that Binds Membranes.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
5FVL
 
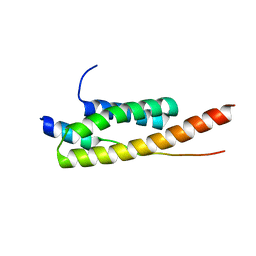 | | Crystal structure of Vps4-Vps20 complex from S.cerevisiae | | Descriptor: | VACUOLAR PROTEIN SORTING-ASSOCIATED PROTEIN 20, VACUOLAR PROTEIN SORTING-ASSOCIATED PROTEIN 4 | | Authors: | Kojima, R, Obita, T, Onoue, K, Mizuguchi, M. | | Deposit date: | 2016-02-09 | | Release date: | 2016-04-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.973 Å) | | Cite: | Structural Fine-Tuning of Mit Interacting Motif 2 (Mim2) and Allosteric Regulation of Escrt-III by Vps4 in Yeast.
J.Mol.Biol., 428, 2016
|
|
5FVK
 
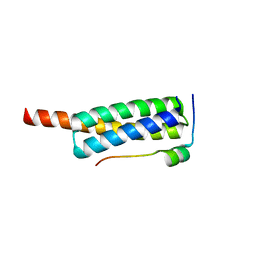 | | Crystal structure of Vps4-Vfa1 complex from S.cerevisiae at 1.66 A resolution. | | Descriptor: | VACUOLAR PROTEIN SORTING-ASSOCIATED PROTEIN 4, VPS4-ASSOCIATED PROTEIN 1 | | Authors: | Kojima, R, Obita, T, Onoue, K, Mizuguchi, M. | | Deposit date: | 2016-02-09 | | Release date: | 2016-04-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.658 Å) | | Cite: | Structural Fine-Tuning of Mit Interacting Motif 2 (Mim2) and Allosteric Regulation of Escrt-III by Vps4 in Yeast.
J.Mol.Biol., 428, 2016
|
|
2XVC
 
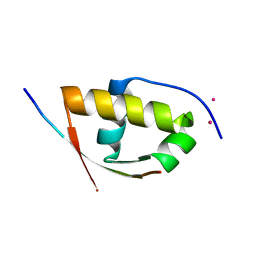 | | Molecular and structural basis of ESCRT-III recruitment to membranes during archaeal cell division | | Descriptor: | CADMIUM ION, CDVA, SSO0911, ... | | Authors: | Samson, R.Y, Obita, T, Hodgson, B, Shaw, M.K, Chong, P.L, Williams, R.L, Bell, S.D. | | Deposit date: | 2010-10-25 | | Release date: | 2011-02-02 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Molecular and Structural Basis of Escrt-III Recruitment to Membranes During Archaeal Cell Division.
Mol.Cell, 41, 2011
|
|
2YMB
 
 | | Structures of MITD1 | | Descriptor: | CHARGED MULTIVESICULAR BODY PROTEIN 1A, MIT DOMAIN-CONTAINING PROTEIN 1 | | Authors: | Hadders, M.A, Agromayor, M, Obita, T, Perisic, O, Caballe, A, Kloc, M, Lamers, M.H, Williams, R.L, Martin-Serrano, J. | | Deposit date: | 2012-10-08 | | Release date: | 2012-10-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.404 Å) | | Cite: | Escrt-III Binding Protein Mitd1 is Involved in Cytokinesis and Has an Unanticipated Pld Fold that Binds Membranes.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
2ZAG
 
 | |
2ZAI
 
 | | Crystal structure of the soluble domain of STT3 from P. furiosus | | Descriptor: | CALCIUM ION, CHLORIDE ION, Oligosaccharyl transferase stt3 subunit related protein | | Authors: | Maita, N. | | Deposit date: | 2007-10-05 | | Release date: | 2007-12-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure-guided identification of a new catalytic motif of oligosaccharyltransferase
Embo J., 27, 2008
|
|
