8RXU
 
 | | Crystal structure of octaheme nitrite reductase from Trichlorobacter ammonificans in space group P21 | | Descriptor: | CALCIUM ION, HEME C, Octaheme nitrite reductase, ... | | Authors: | Polyakov, K.M, Safonova, T.N, Osipov, E, Popov, A.N, Tikhonova, T.V, Popov, V.O. | | Deposit date: | 2024-02-08 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.737 Å) | | Cite: | Crystal structure of octaheme nitrite reductase from Trichlorobacter ammonificans in space group P21
To Be Published
|
|
8RV0
 
 | | Crystal structure of octaheme nitrite reductase from Trichlorobacter ammonificans in complex with nitrite | | Descriptor: | CALCIUM ION, HEME C, NITRIC OXIDE, ... | | Authors: | Polyakov, K.M, Safonova, T.N, Osipov, E, Popov, A.N, Tikhonova, T.V, Popov, V.O. | | Deposit date: | 2024-01-31 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal structure of octaheme nitrite reductase from Trichlorobacter ammonificans in complex with nitrite
To Be Published
|
|
8RVM
 
 | | Crystal structure of octaheme nitrite reductase from Trichlorobacter ammonificans in space group P63 | | Descriptor: | CALCIUM ION, HEME C, Octaheme nitrite c cytochrome c reductase, ... | | Authors: | Polyakov, K.M, Safoonova, T.N, Osipov, E, Popov, A.N, Tikhonova, T.V, Popov, V.O. | | Deposit date: | 2024-02-01 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of octaheme nitrite reductase from Trichlorobacter ammonificans in space group P63
To Be Published
|
|
3SQR
 
 | | Crystal structure of laccase from Botrytis aclada at 1.67 A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, COPPER (II) ION, SULFATE ION, ... | | Authors: | Osipov, E.M, Polyakov, K.M, Tikhonova, T.V, Dorovatovsky, P.V, Ludwig, R, Kittl, R, Shleev, S.V, Popov, V.O. | | Deposit date: | 2011-07-06 | | Release date: | 2012-07-11 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Effect of the L499M mutation of the ascomycetous Botrytis aclada laccase on redox potential and catalytic properties.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3V9E
 
 | | Structure of the L499M mutant of the laccase from B.aclada | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, COPPER (II) ION, GLYCEROL, ... | | Authors: | Osipov, E.M, Polyakov, K.M, Tikhonova, T.V, Dorovatovsky, P.V, Ludwig, R, Kittl, R, Shleev, S.V, Popov, V.O. | | Deposit date: | 2011-12-27 | | Release date: | 2013-01-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Effect of the L499M mutation of the ascomycetous Botrytis aclada laccase on redox potential and catalytic properties.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
6SJI
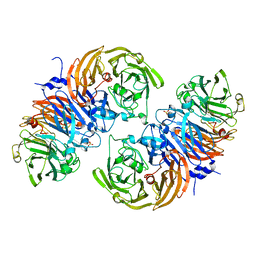 
 | | The structure of thiocyanate dehydrogenase from Thioalkalivibrio paradoxus mutant with His 482 replaced by Gln | | Descriptor: | COPPER (II) ION, SULFATE ION, thiocyanate dehydrogenase | | Authors: | Polyakov, K.M, Tikhonova, T.V, Rakitina, T.V, Osipov, E, Popov, V.O. | | Deposit date: | 2019-08-13 | | Release date: | 2019-09-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Trinuclear copper biocatalytic center forms an active site of thiocyanate dehydrogenase.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6UWE
 
 | | Crystal structure of recombinant thiocyanate dehydrogenase from Thioalkalivibrio paradoxus saturated with copper | | Descriptor: | COPPER (II) ION, UNKNOWN ATOM OR ION, thiocyanate dehydrogenase | | Authors: | Shabalin, I.G, Osipov, E, Tikhonova, T.V, Rakitina, T.V, Boyko, K.M, Popov, V.O. | | Deposit date: | 2019-11-05 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Trinuclear copper biocatalytic center forms an active site of thiocyanate dehydrogenase.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
