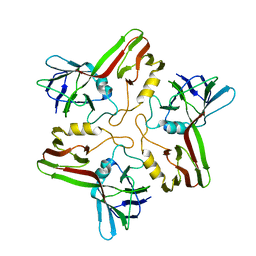
8A02
 
 | |
7ZZZ
 
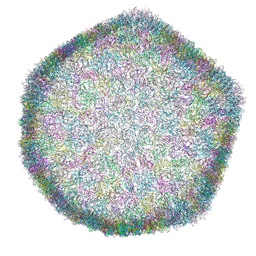 | | Bacteriophage phiCjT23 capsid | | Descriptor: | Major capsid protein P5, Spike protein P13 N-terminal, capsid internal domain, ... | | Authors: | Rissanen, I, Kejzar, N, Huiskonen, J.T. | | Deposit date: | 2022-05-26 | | Release date: | 2022-12-14 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Cryo-EM structure of ssDNA bacteriophage Phi CjT23 provides insight into early virus evolution.
Nat Commun, 13, 2022
|
|
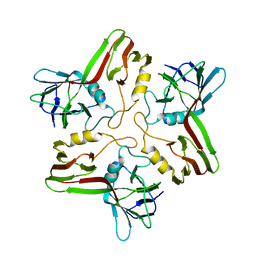
8A01
 
 | |
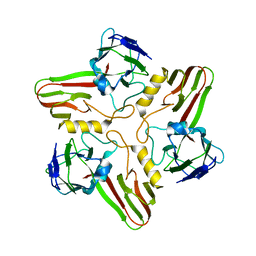
8A04
 
 | |
8A03
 
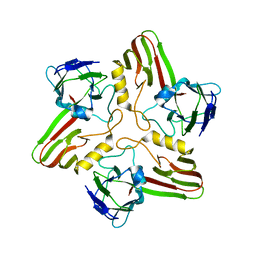 | |
8A05
 
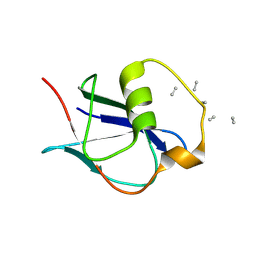 | | Bacteriophage phiCjT23 spike protein penton domain | | Descriptor: | Spike protein P13 N-terminal, capsid internal domain, Unknown vertex protein | | Authors: | Rissanen, I, Huiskonen, J.T. | | Deposit date: | 2022-05-26 | | Release date: | 2022-12-14 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Cryo-EM structure of ssDNA bacteriophage Phi CjT23 provides insight into early virus evolution.
Nat Commun, 13, 2022
|
|
8A06
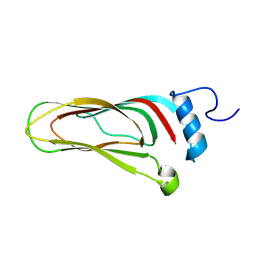 
 | |
9B64
 | |
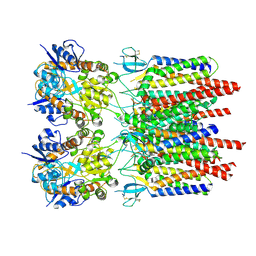
9B6A
 
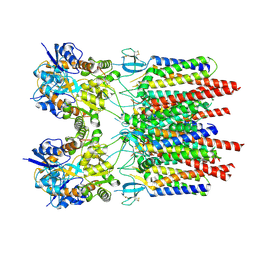 | |
9B69
 
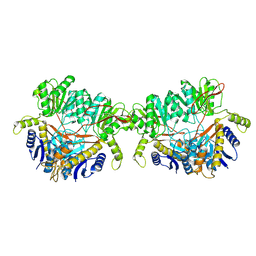 | | GluA2 flip Q in complex with TARPgamma2 at pH8, class12, structure of NTD | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Isoform Flip of Glutamate receptor 2, ... | | Authors: | Nakagawa, T, Greger, I.H. | | Deposit date: | 2024-03-23 | | Release date: | 2024-07-31 | | Method: | ELECTRON MICROSCOPY (3.69 Å) | | Cite: | Proton-triggered rearrangement of the AMPA receptor N-terminal domains impacts receptor kinetics and synaptic localization
To Be Published
|
|
9B68
 
 | | GluA2 flip Q in complex with TARPgamma2 at pH8, class1, structure of NTD | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Isoform Flip of Glutamate receptor 2, ... | | Authors: | Nakagawa, T, Greger, I.H. | | Deposit date: | 2024-03-23 | | Release date: | 2024-07-31 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Proton-triggered rearrangement of the AMPA receptor N-terminal domains impacts receptor kinetics and synaptic localization
To Be Published
|
|
9B5Z
 | |
9B63
 | |
9B60
 
 | |
9B61
 
 | |
9B67
 | |
