2UWJ
 
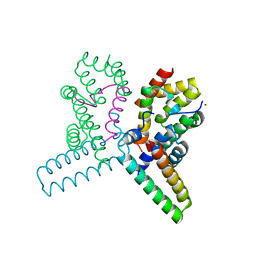 | | Structure of the heterotrimeric complex which regulates type III secretion needle formation | | Descriptor: | NICKEL (II) ION, TYPE III EXPORT PROTEIN PSCE, TYPE III EXPORT PROTEIN PSCF, ... | | Authors: | Quinaud, M, Ple, S, Job, V, Contreras-Martel, C, Simorre, J.P, Attree, I, Dessen, A. | | Deposit date: | 2007-03-22 | | Release date: | 2007-05-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the heterotrimeric complex that regulates type III secretion needle formation.
Proc. Natl. Acad. Sci. U.S.A., 104, 2007
|
|
5J76
 
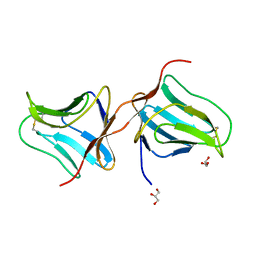 | | Structure of Lectin from Colocasia esculenta(L.) Schott | | Descriptor: | 12kD storage protein, GLYCEROL | | Authors: | Vajravijayan, S, Pletnev, S, Nandhagopal, N, Gunasekaran, K. | | Deposit date: | 2016-04-06 | | Release date: | 2016-06-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural analysis of beta-prism lectin from Colocasia esculenta (L.) S chott.
Int.J.Biol.Macromol., 91, 2016
|
|
5YCZ
 
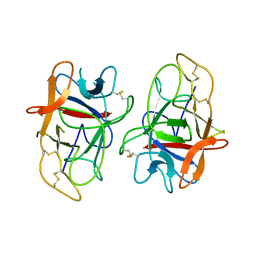 | | Crystal structure of Alocasin, protease inhibitor from Giant Taro (Arum macrorrhizon) | | Descriptor: | Trypsin/chymotrypsin inhibitor | | Authors: | Vajravijayan, S, Pletnev, S, Nandhagopal, N, Gunasekaran, K. | | Deposit date: | 2017-09-08 | | Release date: | 2018-06-13 | | Last modified: | 2018-11-28 | | Method: | X-RAY DIFFRACTION (2.502 Å) | | Cite: | Crystal structure of a novel Kunitz type inhibitor, alocasin with anti-Aedes aegypti activity targeting midgut proteases.
Pest Manag. Sci., 74, 2018
|
|
4ZFS
 
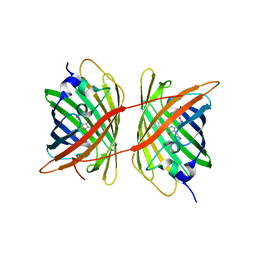 | |
6MAS
 
 | | X-ray Structure of Branchiostoma floridae fluorescent protein lanFP10G | | Descriptor: | GLYCEROL, Uncharacterized protein | | Authors: | Muslinkina, L, Pletneva, N, Pletnev, V, Pletnev, S. | | Deposit date: | 2018-08-28 | | Release date: | 2019-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structural Factors Enabling Successful GFP-Like Proteins with Alanine as the Third Chromophore-Forming Residue.
J. Mol. Biol., 431, 2019
|
|
6M9Y
 | |
6M9X
 
 | |
4HE4
 
 | |
6UBO
 
 | | Fluorogen Activating Protein Dib1 | | Descriptor: | 12-(diethylamino)-2,2-bis(fluoranyl)-4,5-dimethyl-5-aza-3-azonia-2-boranuidatricyclo[7.4.0.0^{3,7}]trideca-1(13),3,7,9,11-pentaen-6-one, CITRIC ACID, Outer membrane lipoprotein Blc, ... | | Authors: | Muslinkina, L, Pletneva, N, Pletnev, V.Z, Pletnev, S. | | Deposit date: | 2019-09-12 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Structure-Based Rational Design of Two Enhanced Bacterial Lipocalin Blc Tags for Protein-PAINT Super-resolution Microscopy.
Acs Chem.Biol., 15, 2020
|
|
6H0I
 
 | | Solution structure of Melampsora larici-populina MlpP4.1 | | Descriptor: | Secreted protein | | Authors: | Tsan, P, Petre, B, Hecker, A, Rouhier, N, Duplessis, S. | | Deposit date: | 2018-07-09 | | Release date: | 2018-08-22 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural genomics applied to the rust fungus Melampsora larici-populina reveals two candidate effector proteins adopting cystine knot and NTF2-like protein folds.
Sci Rep, 9, 2019
|
|
8EEU
 | | Venezuelan equine encephalitis virus-like particle in complex with Fab SKT05 | | Descriptor: | Coat protein, Fab SKT05 heavy chain, Fab SKT05 light chain | | Authors: | Tsybovsky, Y, Pletnev, S, Verardi, R, Roederer, M, Kwong, P.D. | | Deposit date: | 2022-09-07 | | Release date: | 2023-07-19 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Vaccine elicitation and structural basis for antibody protection against alphaviruses.
Cell, 186, 2023
|
|
8EEV
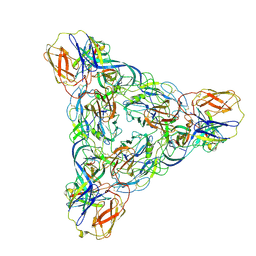 
 | | Venezuelan equine encephalitis virus-like particle in complex with Fab SKT-20 | | Descriptor: | Coat protein, Fab SKT20 heavy chain, Fab SKT20 light chain | | Authors: | Tsybovsky, Y, Pletnev, S, Verardi, R, Roederer, M, Kwong, P.D. | | Deposit date: | 2022-09-07 | | Release date: | 2023-07-19 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Vaccine elicitation and structural basis for antibody protection against alphaviruses.
Cell, 186, 2023
|
|
4ORN
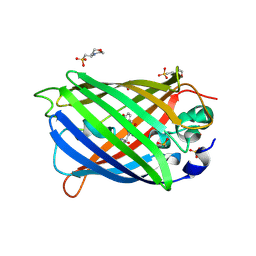 
 | | Blue Fluorescent Protein mKalama1 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CHLORIDE ION, SULFATE ION, ... | | Authors: | Vegh, R.B, Bloch, D.A, Bommarius, A.S, Verkhovsky, M, Pletnev, S, Iwai, H, Bochenkova, A.V, Solntsev, K.M. | | Deposit date: | 2014-02-11 | | Release date: | 2015-02-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Hidden photoinduced reactivity of the blue fluorescent protein mKalama1.
Phys Chem Chem Phys, 17, 2015
|
|
6KBQ
 
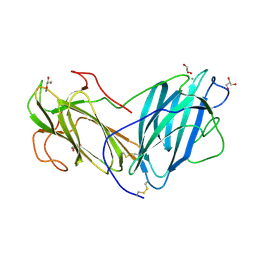 | | Crystal Structure of Lectin from Pleurotus ostreatus in complex with Glycerol | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, GLYCEROL, ... | | Authors: | Vajravijayan, S, Pletnev, S, Luo, Z, Gunasekaran, K, Nandhagopal, N. | | Deposit date: | 2019-06-26 | | Release date: | 2020-08-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.299 Å) | | Cite: | Crystallographic and calorimetric analysis on Pleurotus ostreatus lectin and its sugar complexes - promiscuous binding driven by geometry.
Int.J.Biol.Macromol., 152, 2020
|
|
6KC2
 
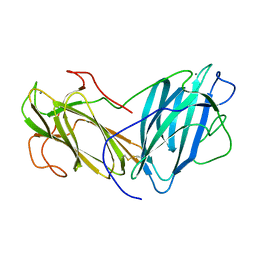 | | Crystal Structure of Lectin from Pleurotus ostreatus in complex with Rhamnose | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Lectin, ... | | Authors: | Gunasekaran, K, Pletnev, S, Luo, Z, Vajravijayan, S, Nandhagopal, N. | | Deposit date: | 2019-06-26 | | Release date: | 2020-08-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.246 Å) | | Cite: | Crystallographic and calorimetric analysis on Pleurotus ostreatus lectin and its sugar complexes - promiscuous binding driven by geometry.
Int.J.Biol.Macromol., 152, 2020
|
|
6KBJ
 
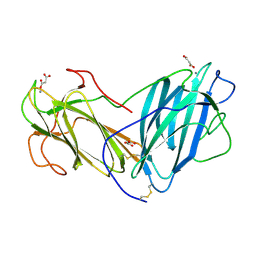 | | Structure of Lectin from Pleurotus ostreatus in complex with malonate | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Lectin, ... | | Authors: | Vajravijayan, S, Pletnev, S, Luo, Z, Gunasekaran, K, Nandhagopal, N. | | Deposit date: | 2019-06-25 | | Release date: | 2020-08-12 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystallographic and calorimetric analysis on Pleurotus ostreatus lectin and its sugar complexes - promiscuous binding driven by geometry.
Int.J.Biol.Macromol., 152, 2020
|
|
3F2L
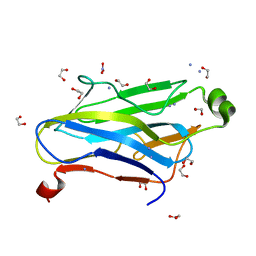 
 | | Crystal structure analysis of the K171A mutation of N-terminal type II cohesin 1 from the cellulosomal ScaB subunit of Acetivibrio cellulolyticus | | Descriptor: | 1,2-ETHANEDIOL, AMMONIUM ION, Cellulosomal scaffoldin adaptor protein B, ... | | Authors: | Frolow, F, Freeman, A, Wine, Y, Eppel, A, Shanzer, M, Stempler, S. | | Deposit date: | 2008-10-30 | | Release date: | 2008-12-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure analysis of the K171A mutation of N-terminal type II cohesin 1 from the cellulosomal ScaB subunit of Acetivibrio cellulolyticus
To be Published
|
|
4N3D
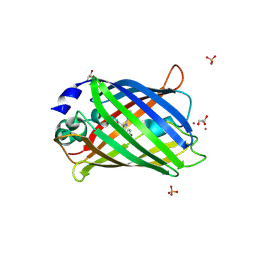 
 | | Crystal structure of the dimeric variant EGFP-K162Q in P61 space group | | Descriptor: | GLYCEROL, Green fluorescent protein, PHOSPHATE ION, ... | | Authors: | Pletneva, N.V, Pletnev, V.Z, Pletnev, S.V. | | Deposit date: | 2013-10-07 | | Release date: | 2014-08-27 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Three dimensional structure of the dimeric gene-engineered variant of green fluorescent protein egfp-K162Q in P61 crystal space group
Rus.J.Bioorg.Chem., 40, 2014
|
|
2PXW
 | | Crystal Structure of N66D Mutant of Green Fluorescent Protein from Zoanthus sp. at 2.4 A Resolution (Transition State) | | Descriptor: | GFP-like fluorescent chromoprotein FP506 | | Authors: | Pletnev, S.V, Pletneva, N.V, Tikhonova, T.V, Pletnev, V.Z. | | Deposit date: | 2007-05-14 | | Release date: | 2007-09-25 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Refined crystal structures of red and green fluorescent proteins from the button polyp Zoanthus.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
5HBJ
 
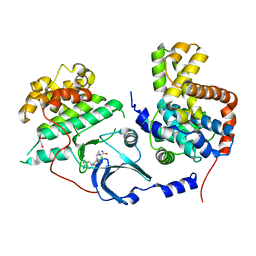 | | CDK8-CYCC IN COMPLEX WITH 8-[2-Amino-3-chloro-5-(1-methyl-1H-indazol-5-yl)-pyridin-4-yl]-2,8-diaza-spiro[4.5]decan-1-one | | Descriptor: | 1,2-ETHANEDIOL, 8-[2-azanyl-3-chloranyl-5-(1-methylindazol-5-yl)pyridin-4-yl]-2,8-diazaspiro[4.5]decan-1-one, Cyclin-C, ... | | Authors: | Musil, D, Blagg, J, Mallinger, A. | | Deposit date: | 2015-12-31 | | Release date: | 2016-02-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Discovery of Potent, Selective, and Orally Bioavailable Small-Molecule Modulators of the Mediator Complex-Associated Kinases CDK8 and CDK19.
J.Med.Chem., 59, 2016
|
|
4RYW
 | |
4ZBL
 | |
5WQU
 
 | | Crystal structure of Sweet Potato Beta-Amylase complexed with Maltotetraose | | Descriptor: | Beta-amylase, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Vajravijayan, S, Sergei, P, Nandhagopal, N, Gunasekaran, K. | | Deposit date: | 2016-11-28 | | Release date: | 2017-12-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Structural insights on starch hydrolysis by plant beta-amylase and its evolutionary relationship with bacterial enzymes
Int. J. Biol. Macromol., 113, 2018
|
|
5WQS
 
 | | Crystal structure of Apo Beta-Amylase from Sweet potato | | Descriptor: | Beta-amylase, ISOPROPYL ALCOHOL | | Authors: | Vajravijayan, S, Sergei, P, Nandhagopal, N, Gunasekaran, K. | | Deposit date: | 2016-11-28 | | Release date: | 2017-12-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural insights on starch hydrolysis by plant beta-amylase and its evolutionary relationship with bacterial enzymes
Int. J. Biol. Macromol., 113, 2018
|
|
6ZQO
 | | EYFP mutant - F165G | | Descriptor: | G protein/GFP fusion protein, SULFATE ION | | Authors: | Pletnev, V.Z, Pletnev, S.V, Pletneva, N.V. | | Deposit date: | 2020-07-10 | | Release date: | 2021-06-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Amino acid residue at the 165th position tunes EYFP chromophore maturation. A structure-based design.
Comput Struct Biotechnol J, 19, 2021
|
|
