5L83
 
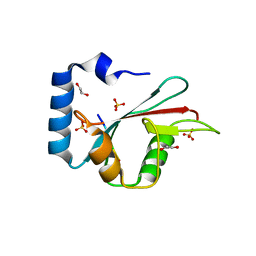 | | Complex of potato ATG8 protein with a peptide from Irish potato famine pathogen effector protein PexRD54 | | Descriptor: | 1,2-ETHANEDIOL, ASP-TRP-GLU-ILE-VAL, Autophagy-related protein, ... | | Authors: | Maqbool, A, Hughes, R.K, Banfield, M.J. | | Deposit date: | 2016-06-06 | | Release date: | 2016-08-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis of Host Autophagy-related Protein 8 (ATG8) Binding by the Irish Potato Famine Pathogen Effector Protein PexRD54.
J.Biol.Chem., 291, 2016
|
|
1ZWX
 
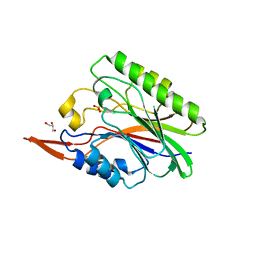 | | Crystal Structure of SmcL | | Descriptor: | GLYCEROL, PHOSPHATE ION, sphingomyelinase-c | | Authors: | Openshaw, A.E.A, Race, P.R, Monzo, H.J, Vasquez-Boland, J.A, Banfield, M.J. | | Deposit date: | 2005-06-06 | | Release date: | 2005-08-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of SmcL, a bacterial neutral sphingomyelinase C from Listeria.
J.Biol.Chem., 280, 2005
|
|
1WKO
 
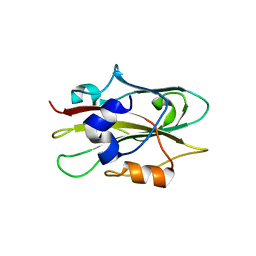 | |
3GQM
 
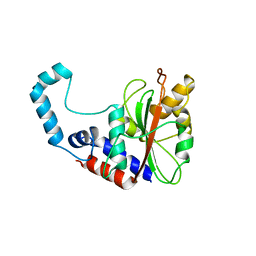 | |
3GQJ
 
 | |
3ZRG
 
 | | Crystal structure of RxLR effector PexRD2 from Phytophthora infestans | | Descriptor: | BROMIDE ION, PEXRD2 FAMILY SECRETED RXLR EFFECTOR PEPTIDE, PUTATIVE | | Authors: | King, S.R.F, Boutemy, L.S, Win, J, Hughes, R.K, Clarke, T.A, Blumenschein, T.M.A, Kamoun, S, Banfield, M.J. | | Deposit date: | 2011-06-16 | | Release date: | 2011-08-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structures of Phytophthora Rxlr Effector Proteins: A Conserved But Adaptable Fold Underpins Functional Diversity.
J.Biol.Chem., 286, 2011
|
|
3ZGK
 
 | | NMR solution structure of the RXLR effector AVR3a11 from Phytophthora Capsici | | Descriptor: | AVR3A11 | | Authors: | Tolchard, J, Chambers, V.S, Boutemy, L.S, Gathercole, R.L, Banfield, M.J, Blumenschein, T.M. | | Deposit date: | 2012-12-18 | | Release date: | 2014-01-08 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of the Avr3A11 from Phytophthora Capsi
To be Published
|
|
3ZR8
 
 | | Crystal structure of RxLR effector Avr3a11 from Phytophthora capsici | | Descriptor: | AVR3A11, CHLORIDE ION, TRIETHYLENE GLYCOL | | Authors: | Boutemy, L.S, King, S.R.F, Win, J, Hughes, R.K, Clarke, T.A, Blumenschein, T.M.A, Kamoun, S, Banfield, M.J. | | Deposit date: | 2011-06-15 | | Release date: | 2011-08-03 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (0.9 Å) | | Cite: | Structures of Phytophthora Rxlr Effector Proteins: A Conserved But Adaptable Fold Underpins Functional Diversity.
J.Biol.Chem., 286, 2011
|
|
4B6X
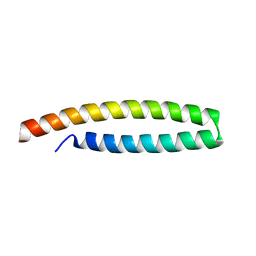 
 | | Crystal structure of in planta processed AvrRps4 | | Descriptor: | ACETATE ION, AVIRULENCE PROTEIN | | Authors: | Sohn, K.H, Hughes, R.K, Piquerez, S.J, Jones, J.D.G, Banfield, M.J. | | Deposit date: | 2012-08-15 | | Release date: | 2012-09-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Distinct Regions of the Pseudomonas Syringae Coiled-Coil Effector Avrrps4 are Required for Activation of Immunity.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
6R8M
 
 | | Complex of rice blast (Magnaporthe oryzae) effector protein AVR-PikE with an engineered HMA domain of Pikp-1 from rice (Oryza sativa) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, AVR-Pik protein, CHLORIDE ION, ... | | Authors: | De la Concepcion, J.C, Franceschetti, M, Banfield, M.J. | | Deposit date: | 2019-04-02 | | Release date: | 2019-04-24 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Protein engineering expands the effector recognition profile of a rice NLR immune receptor.
Elife, 8, 2019
|
|
6S2P
 
 | |
6R8K
 
 | |
1NNE
 
 | | Crystal Structure of the MutS-ADPBeF3-DNA complex | | Descriptor: | 1,2-ETHANEDIOL, 5'-D(*GP*CP*GP*AP*CP*GP*CP*TP*AP*GP*CP*GP*TP*GP*CP*GP*GP*CP*TP*CP*GP*TP*C)-3', 5'-D(P*GP*GP*AP*CP*GP*AP*GP*CP*CP*GP*CP*CP*GP*CP*TP*AP*GP*CP*GP*TP*CP*G)-3', ... | | Authors: | Alani, E, Lee, J.Y, Schofield, M.J, Kijas, A.W, Hsieh, P, Yang, W. | | Deposit date: | 2003-01-13 | | Release date: | 2003-05-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Crystal structure and biochemical analysis of the MutS-ADP-Beryllium Fluoride complex
suggests a conserved mechanism for ATP interactions in mismatch repair
J.Biol.Chem., 278, 2003
|
|
4BUG
 
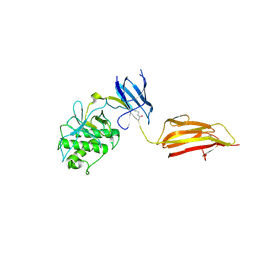 | | Pilus-presented adhesin, Spy0125 (Cpa), Cys426Ala mutant | | Descriptor: | ANCILLARY PROTEIN 1 | | Authors: | Walden, M, Crow, A, Nelson, M, Banfield, M.J. | | Deposit date: | 2013-06-20 | | Release date: | 2013-10-09 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Intramolecular Isopeptide But not Internal Thioester Bonds Confer Proteolytic and Significant Thermal Stability to the S. Pyogenes Pilus Adhesin Spy0125.
Proteins, 82, 2014
|
|
4FBJ
 
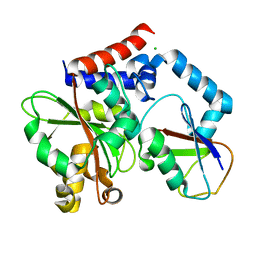 | |
5OD4
 
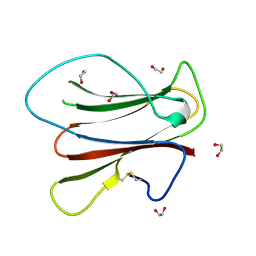 | |
5OET
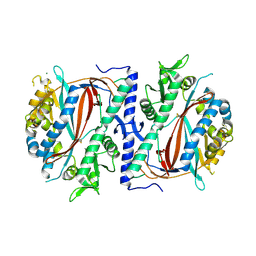 
 | | The structure of a glutathione synthetase like-effector (GSS30) from Globodera pallida in apoform. | | Descriptor: | Glutathione synthetase-like effector 30 (Gpa-GSS30-apo), MAGNESIUM ION | | Authors: | Lilley, C.J, Maqbool, A, Wu, D, Yusup, H.B, Jones, L.M, Birch, P.R.J, Banfield, M.J, Urwin, P.E, Eves-van den Akker, S. | | Deposit date: | 2017-07-10 | | Release date: | 2018-04-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Effector gene birth in plant parasitic nematodes: Neofunctionalization of a housekeeping glutathione synthetase gene.
PLoS Genet., 14, 2018
|
|
5OEV
 
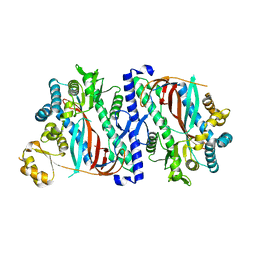 | | The structure of a glutathione synthetase like-effector (GSS22) from Globodera pallida in apoform. | | Descriptor: | Glutathione synthetase-like effector 22 (Gpa-GSS22-apo) | | Authors: | Lilley, C.J, Maqbool, A, Wu, D, Yusup, H.B, Jones, L.M, Birch, P.R.J, Banfield, M.J, Urwin, P.E, Eves-van den Akker, S. | | Deposit date: | 2017-07-10 | | Release date: | 2018-04-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Effector gene birth in plant parasitic nematodes: Neofunctionalization of a housekeeping glutathione synthetase gene.
PLoS Genet., 14, 2018
|
|
5OEU
 
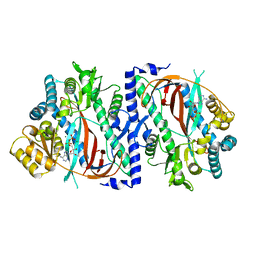 | | The structure of a glutathione synthetase like-effector (GSS22) from Globodera pallida in ADP-bound closed conformation. | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Glutathione synthetase-like effector 22 (Gpa-GSS22-closed), MAGNESIUM ION | | Authors: | Lilley, C.J, Maqbool, A, Wu, D, Yusup, H.B, Jones, L.M, Birch, P.R.J, Banfield, M.J, Urwin, P.E, Eves-van den Akker, S. | | Deposit date: | 2017-07-10 | | Release date: | 2018-04-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Effector gene birth in plant parasitic nematodes: Neofunctionalization of a housekeeping glutathione synthetase gene.
PLoS Genet., 14, 2018
|
|
7NLJ
 
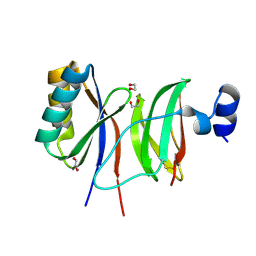 | |
7NMM
 
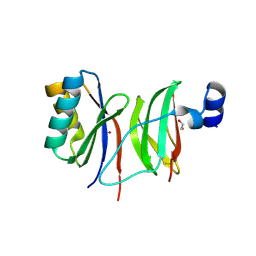 | |
7QPX
 
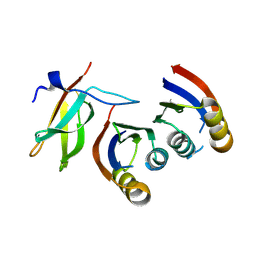 | |
7P8K
 
 | | Crystal structure of in planta processed AvrRps4 in complex with the WRKY domain of RRS1 | | Descriptor: | Avirulence protein,Avirulence protein, Disease resistance protein RRS1, ZINC ION | | Authors: | Mukhi, N, Brown, H, Gorenkin, D, Ding, P, Bentham, A.R, Jones, J.D.G, Banfield, M.J. | | Deposit date: | 2021-07-23 | | Release date: | 2021-08-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Perception of structurally distinct effectors by the integrated WRKY domain of a plant immune receptor.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7QZD
 
 | | Complex of rice blast (Magnaporthe oryzae) effector protein AVR-PikF with an engineered HMA domain of Pikp-1 (Pikp-SNK-EKE) from rice (Oryza sativa) | | Descriptor: | Avr-Pik, Resistance protein Pikp-1 | | Authors: | Maidment, J.H.R, Franceschetti, M, Longya, A, Banfield, M.J. | | Deposit date: | 2022-01-31 | | Release date: | 2022-06-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Effector target-guided engineering of an integrated domain expands the disease resistance profile of a rice NLR immune receptor.
Elife, 12, 2023
|
|
2JKW
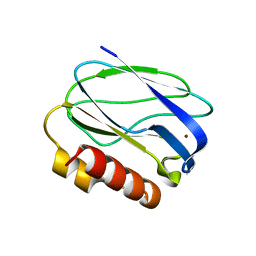 
 | | Pseudoazurin M16F | | Descriptor: | COPPER (II) ION, PSEUDOAZURIN | | Authors: | Yanagisawa, S, Crowley, P.B, Firbank, S.J, Lawler, A.T, Hunter, D.M, McFarlane, W, Li, C, Kohzuma, T, Banfield, M.J, Dennison, C. | | Deposit date: | 2008-09-01 | | Release date: | 2008-11-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Pi-Interaction Tuning of the Active Site Properties of Metalloproteins.
J.Am.Chem.Soc., 130, 2008
|
|
