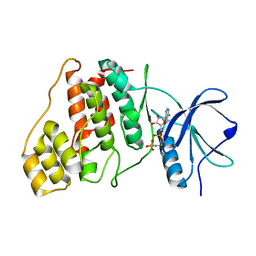
9AV7
 
 | |
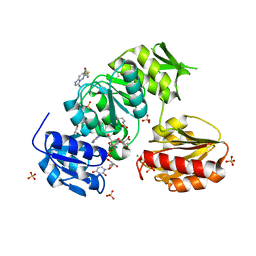
9BN9
 
 | |
9B1Z
 
 | |
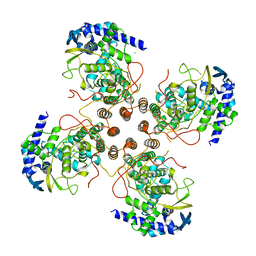
9BJL
 
 | |
9BKY
 
 | |
9AZZ
 
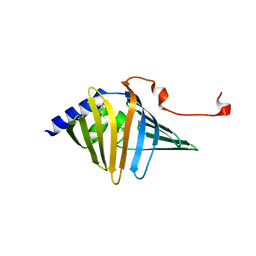 | |
9BKZ
 
 | |
9B21
 
 | | Crystal structure of ADP-ribose diphosphatase from Klebsiella pneumoniae (ADP Ribose bound, Orthorhombic P form) | | Descriptor: | ADP-ribose pyrophosphatase, MAGNESIUM ION, [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL [HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE | | Authors: | Seattle Structural Genomics Center for Infectious Disease, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2024-03-14 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of ADP-ribose diphosphatase from Klebsiella pneumoniae (ADP Ribose bound, Orthorhombic P form)
To be published
|
|
9AV6
 
 | |
9B20
 
 | |
9BKW
 
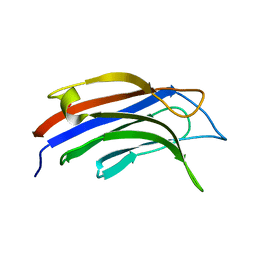 | |
9BN8
 
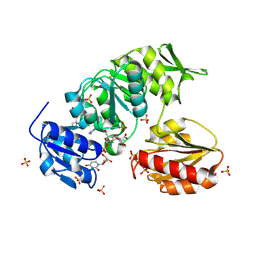 | |
4GL8
 
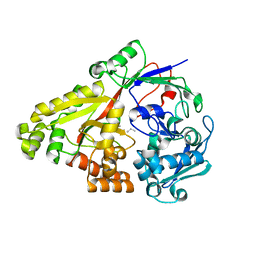 | |
4H51
 
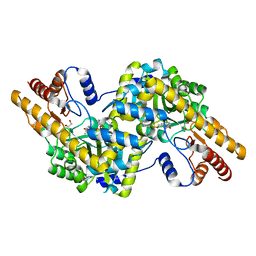 | |
4GRD
 
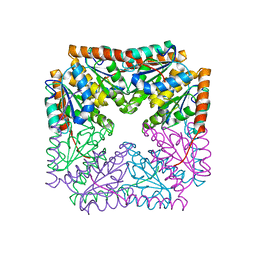 | |
2KN9
 
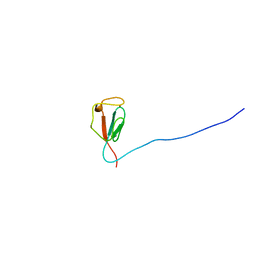 | | Solution structure of zinc-substituted rubredoxin B (Rv3250c) from Mycobacterium tuberculosis. Seattle Structural Genomics Center for Infectious Disease target MytuD.01635.a | | Descriptor: | Rubredoxin, ZINC ION | | Authors: | Buchko, G.W, Hewitt, S.N, Napuli, A.J, Van Voorhis, W.C, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-08-20 | | Release date: | 2009-09-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution-state NMR structure and biophysical characterization of zinc-substituted rubredoxin B (Rv3250c) from Mycobacterium tuberculosis.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
2KOK
 
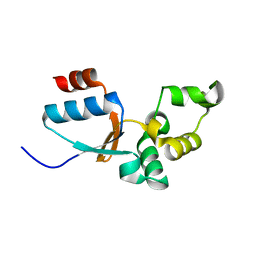 | | Solution structure of an arsenate reductase (ArsC) related protein from Brucella melitensis. Seattle Structural Genomics Center for Infectious Disease target BrabA.00007.a. | | Descriptor: | arsenate reductase | | Authors: | Buchko, G.W, Hewitt, S.N, Napuli, A.J, Van Voorhis, W.C, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-09-23 | | Release date: | 2009-10-13 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution structure of an arsenate reductase-related protein, YffB, from Brucella melitensis, the etiological agent responsible for brucellosis.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
2KE0
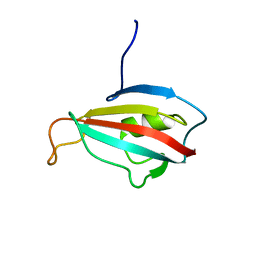 
 | | Solution structure of peptidyl-prolyl cis-trans isomerase from Burkholderia pseudomallei | | Descriptor: | Peptidyl-prolyl cis-trans isomerase | | Authors: | Zheng, S, Leeper, T, Napuli, A, Nakazawa, S.H, Varani, G, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-01-21 | | Release date: | 2009-03-03 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The structure of a Burkholderia pseudomallei immunophilin-inhibitor complex reveals new approaches to antimicrobial development.
Biochem.J., 437, 2011
|
|
4POB
 
 | |
2KHP
 
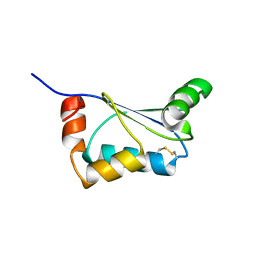 | |
2KDN
 
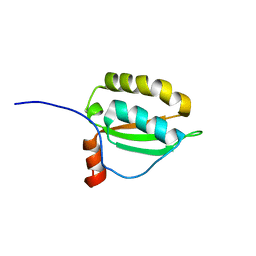 | | Solution structure of PFE0790c, a putative bolA-like protein from the protozoan parasite Plasmodium falciparum. | | Descriptor: | Putative uncharacterized protein PFE0790c | | Authors: | Buchko, G.W, Yee, A, Semesi, A, Hui, R, Arrowsmith, C.H, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-01-12 | | Release date: | 2009-01-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution-state NMR structure of the putative morphogene protein BolA (PFE0790c) from Plasmodium falciparum.
Acta Crystallogr F Struct Biol Commun, 71, 2015
|
|
2KHR
 
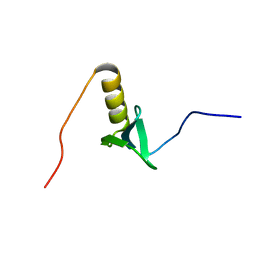 | |
2KO7
 
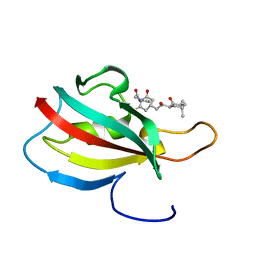 | | Solution structure of peptidyl-prolyl cis-trans isomerase from Burkholderia pseudomallei complexed with Cycloheximide-N-ethylethanoate | | Descriptor: | Peptidyl-prolyl cis-trans isomerase, ethyl (4-{(2R)-2-[(1S,3S,5S)-3,5-dimethyl-2-oxocyclohexyl]-2-hydroxyethyl}-2,6-dioxopiperidin-1-yl)acetate | | Authors: | Zheng, S, Leeper, T, Varani, G, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-09-11 | | Release date: | 2009-09-29 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The structure of a Burkholderia pseudomallei immunophilin-inhibitor complex reveals new approaches to antimicrobial development.
Biochem.J., 437, 2011
|
|
2KZ0
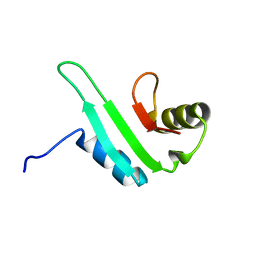 
 | |
2KVC
 
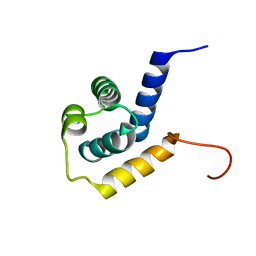 | | Solution structure of the Mycobacterium tuberculosis protein Rv0543c, a member of the DUF3349 superfamily. Seattle Structural Genomics Center for Infectious Disease target MytuD.17112.a | | Descriptor: | Putative uncharacterized protein | | Authors: | Buchko, G.W, Kim, C.Y, Terwilliger, T.C, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2010-03-12 | | Release date: | 2010-03-23 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Inaugural structure from the DUF3349 superfamily of proteins, Mycobacterium tuberculosis Rv0543c.
Arch.Biochem.Biophys., 506, 2011
|
|
