3R6F
 
 | |
3SC4
 
 | |
3SIB
 
 | |
3S6D
 
 | |
3SIA
 
 | |
3S6O
 
 | |
3S5P
 
 | |
3S6L
 | |
3SLG
 
 | |
3S4K
 | |
3SJS
 
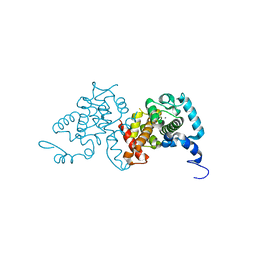 | |
3QRH
 
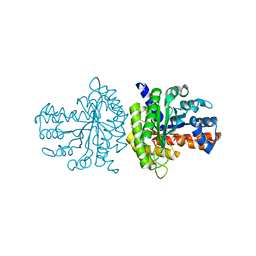 | |
3TE8
 
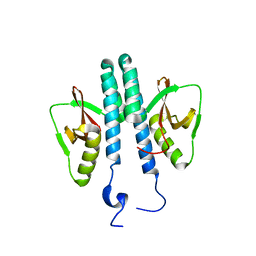 | |
3S99
 
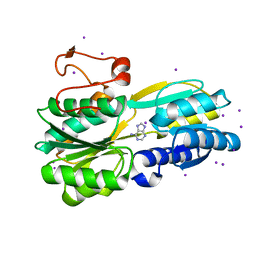 | |
3T80
 
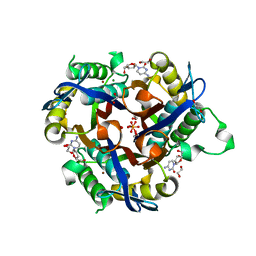 | | Crystal structure of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase from Salmonella typhimurium bound to cytidine | | Descriptor: | 1,2-ETHANEDIOL, 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, 4-AMINO-1-BETA-D-RIBOFURANOSYL-2(1H)-PYRIMIDINONE, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID), Staker, B.L, Edwards, T.E. | | Deposit date: | 2011-07-31 | | Release date: | 2011-09-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase from Salmonella typhimurium bound to cytidine
To be Published
|
|
3RQI
 
 | |
3TK8
 
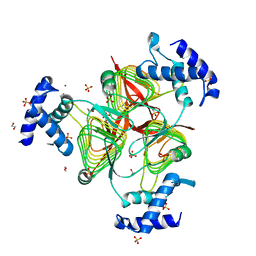 | | Structure of a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase from Burkholderia pseudomallei | | Descriptor: | 1,2-ETHANEDIOL, 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase, SULFATE ION, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-08-25 | | Release date: | 2011-10-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase from Burkholderia pseudomallei
TO BE PUBLISHED
|
|
3U04
 
 | |
3TV2
 
 | |
5KOB
 
 | |
5BQ2
 
 | |
4GNV
 
 | |
4ECP
 
 | |
4EFZ
 
 | |
4EGJ
 
 | |
