1PXQ
 
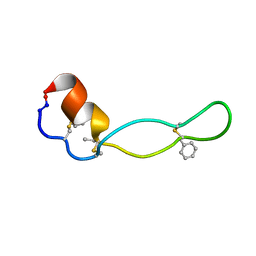 | | Structure of Subtilisin A | | 分子名称: | Subtilisin A | | 著者 | Kawulka, K.E, Sprules, T, McKay, R.T, Mercier, P, Diaper, C.M, Zuber, P, Vederas, J.C. | | 登録日 | 2003-07-04 | | 公開日 | 2004-06-22 | | 最終更新日 | 2011-10-05 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structure of subtilisin A, a cyclic antimicrobial peptide from Bacillus subtilis with unusual sulfur to alpha-carbon cross-links: formation and reduction of alpha-thio-alpha-amino acid derivatives
Biochemistry, 43, 2004
|
|
1Z3E
 
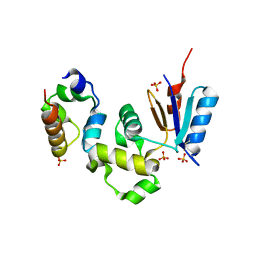 | | Crystal Structure of Spx in Complex with the C-terminal Domain of the RNA Polymerase Alpha Subunit | | 分子名称: | DNA-directed RNA polymerase alpha chain, Regulatory protein spx, SULFATE ION | | 著者 | Newberry, K.J, Nakano, S, Zuber, P, Brennan, R.G. | | 登録日 | 2005-03-11 | | 公開日 | 2005-10-11 | | 最終更新日 | 2017-10-11 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Crystal structure of the Bacillus subtilis anti-alpha, global transcriptional regulator, Spx, in complex with the {alpha} C-terminal domain of RNA polymerase
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
3IHQ
 
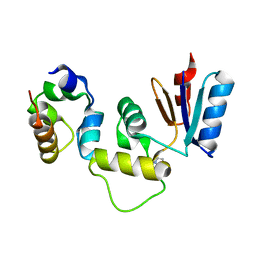 | |
6XE0
 
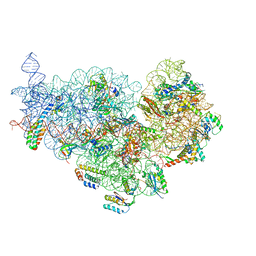 | | Cryo-EM structure of NusG-CTD bound to 70S ribosome (30S: NusG-CTD fragment) | | 分子名称: | 16s rRNA, 30S ribosomal protein S10, 30S ribosomal protein S11, ... | | 著者 | Washburn, R, Zuber, P, Sun, M, Hashem, Y, Shen, B, Li, W, Harvey, S, Acosta-Reyes, F.J, Knauer, S.H, Frank, J, Gottesman, M.E. | | 登録日 | 2020-06-11 | | 公開日 | 2020-07-29 | | 最終更新日 | 2024-03-06 | | 実験手法 | ELECTRON MICROSCOPY (6.8 Å) | | 主引用文献 | Escherichia coli NusG Links the Lead Ribosome with the Transcription Elongation Complex.
Iscience, 23, 2020
|
|
