1VTB
 
 | |
2FPT
 
 | | Dual Binding Mode of a Novel Series of DHODH inhibitors | | 分子名称: | 2-({[3,5-DIFLUORO-3'-(TRIFLUOROMETHOXY)BIPHENYL-4-YL]AMINO}CARBONYL)CYCLOPENT-1-ENE-1-CARBOXYLIC ACID, ACETATE ION, Dihydroorotate dehydrogenase, ... | | 著者 | Baumgartner, R, Walloschek, M, Kralik, M, Gotschlich, A, Tasler, S, Leban, J. | | 登録日 | 2006-01-17 | | 公開日 | 2007-01-23 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Dual binding mode of a novel series of DHODH inhibitors.
J.Med.Chem., 49, 2006
|
|
116D
 
 | |
117D
 
 | |
2IWH
 
 | | Structure of yeast Elongation Factor 3 in complex with ADPNP | | 分子名称: | ELONGATION FACTOR 3A, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, SULFATE ION | | 著者 | Andersen, C.B.F, Becker, T, Blau, M, Anand, M, Halic, M, Balar, B, Mielke, T, Boesen, T, Pedersen, J.S, Spahn, C.M.T, Kinzy, T.G, Andersen, G.R, Beckmann, R. | | 登録日 | 2006-06-30 | | 公開日 | 2006-08-09 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Structure of Eef3 and the Mechanism of Transfer RNA Release from the E-Site.
Nature, 443, 2006
|
|
9G2A
 
 | | Staphylococcus aureus MazF in complex with nanobody 4 | | 分子名称: | 3[N-MORPHOLINO]PROPANE SULFONIC ACID, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Endoribonuclease MazF, ... | | 著者 | Prolic-Kalinsek, M, Zorzini, V, Haesaerts, S, Loris, R. | | 登録日 | 2024-07-10 | | 公開日 | 2024-07-17 | | 実験手法 | X-RAY DIFFRACTION (2.046594 Å) | | 主引用文献 | Staphylococcus aureus MazF in complex with nanobody 4
To Be Published
|
|
2IX3
 
 | | Structure of yeast Elongation Factor 3 | | 分子名称: | ELONGATION FACTOR 3, SULFATE ION | | 著者 | Andersen, C.B.F, Becker, T, Blau, M, Anand, M, Halic, M, Balar, B, Mielke, T, Boesen, T, Pedersen, J.S, Spahn, C.M.T, Kinzy, T.G, Andersen, G.R, Beckmann, R. | | 登録日 | 2006-07-06 | | 公開日 | 2006-08-08 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Structure of Eef3 and the Mechanism of Transfer RNA Release from the E-Site.
Nature, 443, 2006
|
|
2IW3
 
 | | Elongation Factor 3 in complex with ADP | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, ELONGATION FACTOR 3A, SULFATE ION | | 著者 | Andersen, C.B.F, Becker, T, Blau, M, Anand, M, Halic, M, Balar, B, Mielke, T, Boesen, T, Pedersen, J.S, Spahn, C.M.T, Kinzy, T.G, Andersen, G.R, Beckmann, R. | | 登録日 | 2006-06-26 | | 公開日 | 2006-08-17 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Structure of Eef3 and the Mechanism of Transfer RNA Release from the E-Site.
Nature, 443, 2006
|
|
1D78
 
 | | HIGH RESOLUTION REFINEMENT OF THE HEXAGONAL A-DNA OCTAMER D(GTGTACAC) AT 1.4 ANGSTROMS RESOLUTION | | 分子名称: | DNA (5'-D(*GP*TP*GP*TP*AP*CP*AP*C)-3') | | 著者 | Thota, N, Li, X.H, Bingman, C.A, Sundaralingam, M. | | 登録日 | 1992-06-12 | | 公開日 | 1993-04-15 | | 最終更新日 | 2023-03-22 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | High-resolution refinement of the hexagonal A-DNA octamer d(GTGTACAC) at 1.4 A.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
2IX8
 
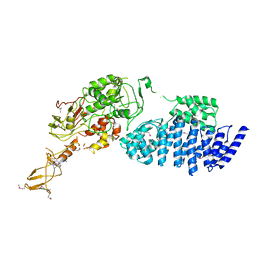 | | MODEL FOR EEF3 BOUND TO AN 80S RIBOSOME | | 分子名称: | ELONGATION FACTOR 3A | | 著者 | Andersen, C.B.F, Becker, T, Blau, M, Anand, M, Halic, M, Balar, B, Mielke, T, Boesen, T, Pedersen, J.S, Spahn, C.M.T, Kinzy, T.G, Andersen, G.R, Beckmann, R. | | 登録日 | 2006-07-07 | | 公開日 | 2007-07-10 | | 最終更新日 | 2017-08-23 | | 実験手法 | ELECTRON MICROSCOPY (6 Å) | | 主引用文献 | Structure of Eef3 and the Mechanism of Transfer RNA Release from the E-Site.
Nature, 443, 2006
|
|
1CEH
 
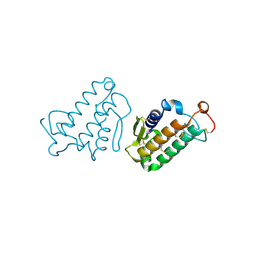 | | STRUCTURE AND FUNCTION OF THE CATALYTIC SITE MUTANT ASP99ASN OF PHOSPHOLIPASE A2: ABSENCE OF CONSERVED STRUCTURAL WATER | | 分子名称: | CALCIUM ION, PHOSPHOLIPASE A2 | | 著者 | Kumar, A, Sekharudu, C, Ramakrishnan, B, Dupureur, C.M, Zhu, H, Tsai, M.-D, Sundaralingam, M. | | 登録日 | 1994-11-16 | | 公開日 | 1995-02-07 | | 最終更新日 | 2024-06-05 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structure and function of the catalytic site mutant Asp 99 Asn of phospholipase A2: absence of the conserved structural water.
Protein Sci., 3, 1994
|
|
1DNS
 
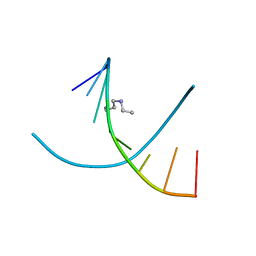 | |
1DN8
 
 | |
1D79
 
 | | HIGH RESOLUTION REFINEMENT OF THE HEXAGONAL A-DNA OCTAMER D(GTGTACAC) AT 1.4 ANGSTROMS RESOLUTION | | 分子名称: | DNA (5'-D(*GP*TP*GP*TP*AP*CP*AP*C)-3') | | 著者 | Thota, N, Li, X.H, Bingman, C.A, Sundaralingam, M. | | 登録日 | 1992-06-12 | | 公開日 | 1993-04-15 | | 最終更新日 | 2023-03-22 | | 実験手法 | X-RAY DIFFRACTION (1.45 Å) | | 主引用文献 | High-resolution refinement of the hexagonal A-DNA octamer d(GTGTACAC) at 1.4 A.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
1J8G
 
 | | X-ray Analysis of a RNA Tetraplex r(uggggu)4 at Ultra-High Resolution | | 分子名称: | 5'-R(*UP*GP*GP*GP*GP*U)-3', CALCIUM ION, SODIUM ION, ... | | 著者 | Deng, J, Xiong, Y, Sundaralingam, M. | | 登録日 | 2001-05-21 | | 公開日 | 2001-11-23 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (0.61 Å) | | 主引用文献 | X-ray analysis of an RNA tetraplex (UGGGGU)(4) with divalent Sr(2+) ions at subatomic resolution (0.61 A).
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1J9H
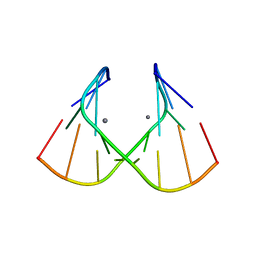 
 | | Crystal Structure of an RNA Duplex with Uridine Bulges | | 分子名称: | 5'-R(*GP*UP*GP*UP*CP*GP*(CBR)P*AP*C)-3', CALCIUM ION | | 著者 | Xiong, Y, Deng, J, Sudarsanakumar, C, Sundaralingam, M. | | 登録日 | 2001-05-25 | | 公開日 | 2001-10-26 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Crystal structure of an RNA duplex r(gugucgcac)(2) with uridine bulges.
J.Mol.Biol., 313, 2001
|
|
1JO2
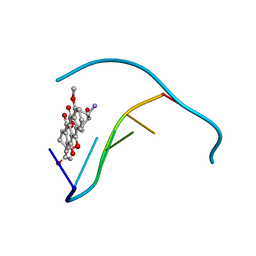 
 | |
8EUI
 
 | | Ytm1 associated nascent 60S ribosome (-fkbp39) State 3 | | 分子名称: | 60S ribosomal protein L10-A, 60S ribosomal protein L11-A, 60S ribosomal protein L13, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-18 | | 公開日 | 2022-11-30 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
8ETI
 
 | | Fkbp39 associated 60S nascent ribosome State 1 | | 分子名称: | 60S ribosomal protein L13, 60S ribosomal protein L14, 60S ribosomal protein L15-A, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-17 | | 公開日 | 2022-11-30 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
8ETH
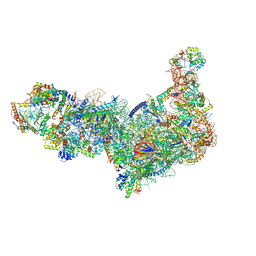 
 | | Ytm1 associated 60S nascent ribosome State 1B | | 分子名称: | 60S ribosomal protein L13, 60S ribosomal protein L14, 60S ribosomal protein L15-A, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-17 | | 公開日 | 2022-11-30 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.8 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
8ETG
 
 | | Fkbp39 associated 60S nascent ribosome State 3 | | 分子名称: | 60S ribosomal protein L13, 60S ribosomal protein L14, 60S ribosomal protein L15-A, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-17 | | 公開日 | 2022-11-30 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
8ESR
 
 | | Ytm1 associated nascent 60S ribosome (-fkbp39) State 2 | | 分子名称: | 25S rRNA (cytosine-C(5))-methyltransferase nop2, 60S ribosomal protein L13, 60S ribosomal protein L14, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-14 | | 公開日 | 2022-11-30 | | 最終更新日 | 2022-12-14 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
8EUP
 
 | | Ytm1 associated 60S nascent ribosome State 1A | | 分子名称: | 60S ribosomal protein L13, 60S ribosomal protein L14, 60S ribosomal protein L15-A, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-19 | | 公開日 | 2022-11-30 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
8EV3
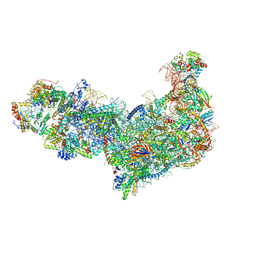 
 | | Ytm1 associated 60S nascent ribosome (-Fkbp39) State 1B | | 分子名称: | 60S ribosomal protein L13, 60S ribosomal protein L14, 60S ribosomal protein L15-A, ... | | 著者 | Zhou, X, Bilokapic, S, Deshmukh, A.A, Halic, M. | | 登録日 | 2022-10-19 | | 公開日 | 2022-11-30 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Chromatin localization of nucleophosmin organizes ribosome biogenesis.
Mol.Cell, 82, 2022
|
|
1RCF
 
 | | STRUCTURE OF THE TRIGONAL FORM OF RECOMBINANT OXIDIZED FLAVODOXIN FROM ANABAENA 7120 AT 1.40 ANGSTROMS RESOLUTION | | 分子名称: | FLAVIN MONONUCLEOTIDE, FLAVODOXIN, SULFATE ION | | 著者 | Burkhart, B, Ramakrishnan, B, Yan, H, Reedstrom, R, Markley, J, Straus, N, Sundaralingam, M. | | 登録日 | 1994-10-31 | | 公開日 | 1995-01-26 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Structure of the trigonal form of recombinant oxidized flavodoxin from Anabaena 7120 at 1.40 A resolution.
Acta Crystallogr.,Sect.D, 51, 1995
|
|
