6U5C
 
 | |
3UG4
 
 | | Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex | | 分子名称: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Alpha-L-arabinofuranosidase, alpha-L-arabinofuranose | | 著者 | Im, D.-H, Miyazaki, K, Wakagi, T, Fushinobu, S. | | 登録日 | 2011-11-02 | | 公開日 | 2012-03-07 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.15 Å) | | 主引用文献 | Crystal Structures of Glycoside Hydrolase Family 51 alpha-L-Arabinofuranosidase from Thermotoga maritima
Biosci.Biotechnol.Biochem., 76, 2012
|
|
3UG5
 
 | | Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex | | 分子名称: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Alpha-L-arabinofuranosidase, beta-D-xylopyranose | | 著者 | Im, D.-H, Miyazaki, K, Wakagi, T, Fushinobu, S. | | 登録日 | 2011-11-02 | | 公開日 | 2012-03-07 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Crystal Structures of Glycoside Hydrolase Family 51 alpha-L-Arabinofuranosidase from Thermotoga maritima
Biosci.Biotechnol.Biochem., 76, 2012
|
|
3UG3
 
 | | Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form | | 分子名称: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Alpha-L-arabinofuranosidase, ... | | 著者 | Im, D.-H, Miyazaki, K, Wakagi, T, Fushinobu, S. | | 登録日 | 2011-11-02 | | 公開日 | 2012-03-07 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Crystal Structures of Glycoside Hydrolase Family 51 alpha-L-Arabinofuranosidase from Thermotoga maritima
Biosci.Biotechnol.Biochem., 76, 2012
|
|
5XFD
 
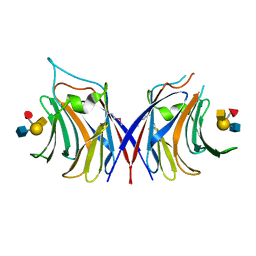 | |
8I2Z
 
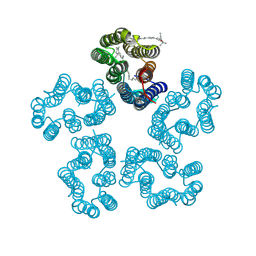 | | Cryo-EM structure of the zeaxanthin-bound kin4B8 | | 分子名称: | RETINAL, Xanthorhodopsin, Zeaxanthin | | 著者 | Murakoshi, S, Chazan, A, Shihoya, W, Beja, O, Nureki, O. | | 登録日 | 2023-01-15 | | 公開日 | 2023-03-29 | | 最終更新日 | 2023-04-19 | | 実験手法 | ELECTRON MICROSCOPY (2 Å) | | 主引用文献 | Phototrophy by antenna-containing rhodopsin pumps in aquatic environments.
Nature, 615, 2023
|
|
5JOO
 
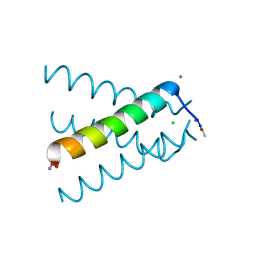 | | XFEL structure of influenza A M2 wild type TM domain at low pH in the lipidic cubic phase at room temperature | | 分子名称: | CALCIUM ION, CHLORIDE ION, Matrix protein 2 | | 著者 | Thomaston, J.L, Woldeyes, R.A, Fraser, J.S, DeGrado, W.F. | | 登録日 | 2016-05-02 | | 公開日 | 2017-08-02 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (1.413 Å) | | 主引用文献 | XFEL structures of the influenza M2 proton channel: Room temperature water networks and insights into proton conduction.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
8XBH
 
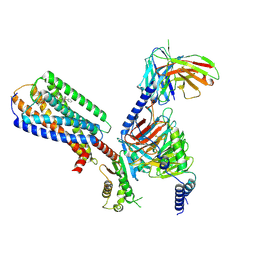 | | Human GPR34 -Gi complex bound to M1 | | 分子名称: | (2~{S})-2-azanyl-3-[[(2~{R})-1-ethoxy-3-[3-[2-[(3-phenoxyphenyl)methoxy]phenyl]propanoyloxy]propan-2-yl]oxy-oxidanyl-phosphoryl]oxy-propanoic acid, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Kawahara, R, Shihoya, W, Nureki, O. | | 登録日 | 2023-12-06 | | 公開日 | 2023-12-27 | | 最終更新日 | 2024-05-15 | | 実験手法 | ELECTRON MICROSCOPY (2.83 Å) | | 主引用文献 | Structural basis for lysophosphatidylserine recognition by GPR34.
Nat Commun, 15, 2024
|
|
8XBI
 
 | | Human GPR34 -Gi complex bound to M1, receptor focused | | 分子名称: | (2~{S})-2-azanyl-3-[[(2~{R})-1-ethoxy-3-[3-[2-[(3-phenoxyphenyl)methoxy]phenyl]propanoyloxy]propan-2-yl]oxy-oxidanyl-phosphoryl]oxy-propanoic acid, Probable G-protein coupled receptor 34 | | 著者 | Kawahara, R, Shihoya, W, Nureki, O. | | 登録日 | 2023-12-06 | | 公開日 | 2023-12-27 | | 最終更新日 | 2024-05-15 | | 実験手法 | ELECTRON MICROSCOPY (3.06 Å) | | 主引用文献 | Structural basis for lysophosphatidylserine recognition by GPR34.
Nat Commun, 15, 2024
|
|
8XBG
 
 | |
8XBE
 
 | | Human GPR34 -Gi complex bound to S3E-LysoPS | | 分子名称: | (2~{S})-2-azanyl-3-[[(2~{R})-1-ethoxy-3-[(~{Z})-octadec-9-enoyl]oxy-propan-2-yl]oxy-oxidanyl-phosphoryl]oxy-propanoic acid, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Kawahara, R, Shihoya, W, Nureki, O. | | 登録日 | 2023-12-06 | | 公開日 | 2024-05-15 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | Structural basis for lysophosphatidylserine recognition by GPR34.
Nat Commun, 15, 2024
|
|
5FVF
 | | Room temperature structure of IrisFP determined by serial femtosecond crystallography. | | 分子名称: | AMMONIUM ION, Green to red photoconvertible GFP-like protein EosFP, SULFATE ION | | 著者 | Colletier, J.P, Gallat, F.X, Coquelle, N, Weik, M. | | 登録日 | 2016-02-06 | | 公開日 | 2016-04-06 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (2.75 Å) | | 主引用文献 | Serial Femtosecond Crystallography and Ultrafast Absorption Spectroscopy of the Photoswitchable Fluorescent Protein Irisfp.
J.Phys.Chem.Lett., 7, 2016
|
|
8CWE
 
 | | 20ns Temperature-Jump (Dark2) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW1
 | | 20us Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW0
 
 | | 20us Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW3
 
 | | 20us Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWB
 
 | | Laser Off Temperature-Jump XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.51 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWC
 
 | | 20ns Temperature-Jump (Light) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVW
 
 | | 20ns Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW5
 
 | | 200us Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVU
 
 | | 20ns Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWF
 
 | | 200us Temperature-Jump (Light) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWD
 
 | | 20ns Temperature-Jump (Dark1) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW6
 
 | | 200us Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWG
 
 | | 200us Temperature-Jump (Dark1) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
