7DXZ
 
 | |
7DXW
 
 | |
7DXR
 
 | |
7DXT
 
 | |
7DYC
 
 | |
7DXV
 
 | |
7DXU
 
 | |
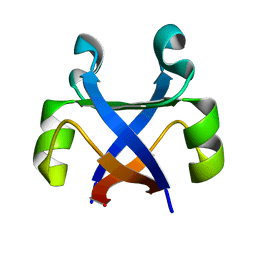
7DXY
 | |
7DXS
 
 | |
5F53
 
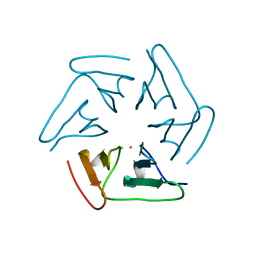 | |
7DU6
 
 | |
7DU7
 
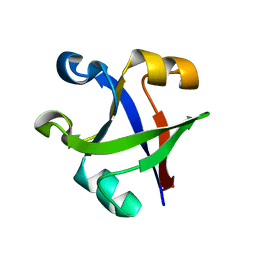 | |
7DVH
 
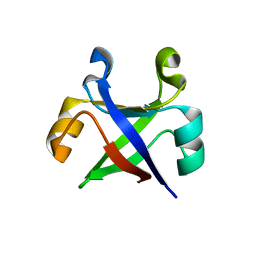 | |
7DVC
 
 | | Crystal structure of the computationally designed reDPBB_sym1 protein | | 分子名称: | ACETATE ION, CHLORIDE ION, reDPBB_sym1 protein | | 著者 | Yagi, S, Tagami, S, Padhi, A.K, Zhang, K.Y.J. | | 登録日 | 2021-01-13 | | 公開日 | 2021-09-29 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.705 Å) | | 主引用文献 | Seven Amino Acid Types Suffice to Create the Core Fold of RNA Polymerase.
J.Am.Chem.Soc., 143, 2021
|
|
7DVF
 
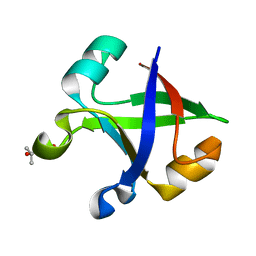 | | Crystal structure of the computationally designed reDPBB_sym2 protein | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, reDPBB_sym2 protein | | 著者 | Yagi, S, Tagami, S, Padhi, A.K, Zhang, K.Y.J. | | 登録日 | 2021-01-13 | | 公開日 | 2021-10-13 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.209 Å) | | 主引用文献 | Seven Amino Acid Types Suffice to Create the Core Fold of RNA Polymerase.
J.Am.Chem.Soc., 143, 2021
|
|
