9G4U
 
 | | Crystal structure of the human METTL3-METTL14 in complex with small molecule inhibitor Compound 31 | | Descriptor: | 4-[(3~{R})-3-(cyclobutylmethylamino)piperidin-1-yl]-1-[(1~{R})-1-[4-(5-methoxypyridin-3-yl)-1,2,3-triazol-1-yl]ethyl]pyridin-2-one, N6-adenosine-methyltransferase catalytic subunit, N6-adenosine-methyltransferase non-catalytic subunit | | Authors: | Dutheuil, G, Oukoloff, K. | | Deposit date: | 2024-07-16 | | Release date: | 2025-02-12 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery, Optimization, and Preclinical Pharmacology of EP652, a METTL3 Inhibitor with Efficacy in Liquid and Solid Tumor Models.
J.Med.Chem., 68, 2025
|
|
9G4W
 
 | | Crystal structure of the human METTL3-METTL14 in complex with small molecule inhibitor Compound 59 | | Descriptor: | 4-[(3~{R})-3-(cyclopropylmethylamino)piperidin-1-yl]-1-[(1~{R})-1-[4-(6-pyrrolidin-1-ylpyrazin-2-yl)-1,2,3-triazol-1-yl]ethyl]pyridin-2-one, CALCIUM ION, N6-adenosine-methyltransferase catalytic subunit, ... | | Authors: | Dutheuil, G, Oukoloff, K. | | Deposit date: | 2024-07-16 | | Release date: | 2025-02-12 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Discovery, Optimization, and Preclinical Pharmacology of EP652, a METTL3 Inhibitor with Efficacy in Liquid and Solid Tumor Models.
J.Med.Chem., 68, 2025
|
|
9GTU
 
 | | Collagen VI alpha 1, 2, 3 heterotrimer recombinant C terminal region. Local refinement. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Collagen alpha-1(VI) chain, ... | | Authors: | Godwin, A, Snee, M, Dajani, R, Becker, M, Roseman, A, Baldock, C. | | Deposit date: | 2024-09-18 | | Release date: | 2025-08-20 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | Collagen VI microfibril structure reveals mechanism for molecular assembly and clustering of inherited pathogenic mutations.
Nat Commun, 16, 2025
|
|
9H0N
 
 | | Fucosylated Lacto-N-biose binding protein from Bifidobacterium longum subsp. infantis in complex with Galacto-N-biose | | Descriptor: | Extracellular solute-binding protein, family 1, SODIUM ION, ... | | Authors: | Jensen, M, Hansen, M.E, Sakanaka, H, Slotboom, D.J, Abou Hachem, M, Morth, J.P. | | Deposit date: | 2024-10-08 | | Release date: | 2025-07-02 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Uptake of fucosylated type I human milk oligosaccharide blocks by Bifidobacterium longum subsp. infantis.
Mbio, 16, 2025
|
|
9H0O
 
 | | Fucosylated Lacto-N-biose binding protein from Bifidobacterium longum subsp. infantis in complex with Lacto-N-biose | | Descriptor: | CALCIUM ION, Extracellular solute-binding protein, family 1, ... | | Authors: | Jensen, M, Hansen, M.E, Sakanaka, H, Slotboom, D.J, Abou Hachem, M, Morth, J.P. | | Deposit date: | 2024-10-08 | | Release date: | 2025-07-02 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Uptake of fucosylated type I human milk oligosaccharide blocks by Bifidobacterium longum subsp. infantis.
Mbio, 16, 2025
|
|
9H0P
 
 | | Fucosylated Lacto-N-biose binding protein from Bifidobacterium longum subsp. infantis in complex with H1 trisaccharide | | Descriptor: | Extracellular solute-binding protein, family 1, alpha-L-fucopyranose-(1-2)-beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-alpha-D-glucopyranose | | Authors: | Jensen, M, Hansen, M.E, Sakanaka, H, Slotboom, D.J, Abou Hachem, M, Morth, J.P. | | Deposit date: | 2024-10-08 | | Release date: | 2025-07-02 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Uptake of fucosylated type I human milk oligosaccharide blocks by Bifidobacterium longum subsp. infantis.
Mbio, 16, 2025
|
|
9H8T
 
 | | Cryo-EM structure of the atovaquone-inhibited Complex III from the Chlorocebus sabaeus respirasome | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, 2-[4-(4-CHLOROPHENYL)CYCLOHEXYLIDENE]-3,4-DIHYDROXY-1(2H)-NAPHTHALENONE, ... | | Authors: | Maclean, A, Muhleip, A. | | Deposit date: | 2024-10-29 | | Release date: | 2025-07-02 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (2.67 Å) | | Cite: | Structure, assembly and inhibition of the Toxoplasma gondii respiratory chain supercomplex.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9HAN
 
 | | Bovine collagen VI local refinement of C-terminal region | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Godwin, A, Snee, M, Roseman, A, Baldock, C. | | Deposit date: | 2024-11-04 | | Release date: | 2025-08-20 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (4.33 Å) | | Cite: | Collagen VI microfibril structure reveals mechanism for molecular assembly and clustering of inherited pathogenic mutations.
Nat Commun, 16, 2025
|
|
9HFL
 
 | | Cryo-EM structure of the human snRNA export complex comprising CBC-PHAX-CRM1-RanGTP and capped-RNA | | Descriptor: | Exportin-1, GTP-binding nuclear protein Ran, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Dubiez, E, Cusack, S, Kadlec, J. | | Deposit date: | 2024-11-18 | | Release date: | 2025-07-16 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (2.62 Å) | | Cite: | Structural basis for the synergistic assembly of the snRNA export complex.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9I4X
 
 | | Toxoplasma gondii cytochrome bc1 complex from the respiratory supercomplex III2-IV inhibited by atovaquone and ELQ-300 | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-[trans-4-(4-chlorophenyl)cyclohexyl]-3-hydroxynaphthalene-1,4-dione, 6-chloranyl-7-methoxy-2-methyl-3-[4-[4-(trifluoromethyloxy)phenoxy]phenyl]-1~{H}-quinolin-4-one, ... | | Authors: | Maclean, A, Muhleip, A. | | Deposit date: | 2025-01-27 | | Release date: | 2025-07-30 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (2.79 Å) | | Cite: | Structure, assembly and inhibition of the Toxoplasma gondii respiratory chain supercomplex.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9IAP
 
 | | Structure of 1 in complex with GDP-KRAS | | Descriptor: | (4~{S})-2-azanyl-4-methyl-4-[3-(3-piperazin-1-ylphenyl)-1,2,4-oxadiazol-5-yl]-6,7-dihydro-5~{H}-1-benzothiophene-3-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J. | | Deposit date: | 2025-02-11 | | Release date: | 2025-08-06 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.182 Å) | | Cite: | Discovery of BI-2493, a Pan-KRAS Inhibitor Showing In Vivo Efficacy.
J.Med.Chem., 68, 2025
|
|
9IAW
 
 | | Structure of 5 in complex with GDP-KRAS | | Descriptor: | (4~{S})-2-azanyl-4-methyl-4-[3-[2-[(2~{S})-2-methylpiperazin-1-yl]pyrimidin-4-yl]-1,2,4-oxadiazol-5-yl]-6,7-dihydro-5~{H}-1-benzothiophene-3-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J. | | Deposit date: | 2025-02-11 | | Release date: | 2025-08-06 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (0.997 Å) | | Cite: | Discovery of BI-2493, a Pan-KRAS Inhibitor Showing In Vivo Efficacy.
J.Med.Chem., 68, 2025
|
|
9IAY
 
 | | Structure of 10 in complex with GDP-KRAS | | Descriptor: | (4S)-2-azanyl-4-methyl-4-[3-[2-[(2S)-2-methyl-1,4-diazepan-1-yl]pyrimidin-4-yl]-1,2,4-oxadiazol-5-yl]-6,7-dihydro-5H-1-benzothiophene-3-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J. | | Deposit date: | 2025-02-11 | | Release date: | 2025-08-06 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (0.953 Å) | | Cite: | Discovery of BI-2493, a Pan-KRAS Inhibitor Showing In Vivo Efficacy.
J.Med.Chem., 68, 2025
|
|
9IB4
 
 | | Structure of 12 in complex with GDP-KRAS | | Descriptor: | (4~{S})-2-azanyl-4-methyl-4-[3-[2-[[(2~{S})-1-methylpyrrolidin-2-yl]methoxy]pyrimidin-4-yl]-1,2,4-oxadiazol-5-yl]-6,7-dihydro-5~{H}-1-benzothiophene-3-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J. | | Deposit date: | 2025-02-11 | | Release date: | 2025-08-06 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.062 Å) | | Cite: | Discovery of BI-2493, a Pan-KRAS Inhibitor Showing In Vivo Efficacy.
J.Med.Chem., 68, 2025
|
|
9IB5
 
 | | Structure of 18 (BI-2493) in complex with GDP-KRAS | | Descriptor: | (7~{S})-2'-azanyl-3-[2-[(2~{S})-2-methylpiperazin-1-yl]pyrimidin-4-yl]spiro[5,6-dihydro-4~{H}-1,2-benzoxazole-7,4'-6,7-dihydro-5~{H}-1-benzothiophene]-3'-carbonitrile, 1,2-ETHANEDIOL, GTPase KRas, ... | | Authors: | Boettcher, J. | | Deposit date: | 2025-02-11 | | Release date: | 2025-08-06 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.011 Å) | | Cite: | Discovery of BI-2493, a Pan-KRAS Inhibitor Showing In Vivo Efficacy.
J.Med.Chem., 68, 2025
|
|
9IIN
 
 | | Structure of CTF18-PCNA with ATP and Mg2+ | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Chromosome transmission fidelity protein 18 homolog, ... | | Authors: | Briola, G.R, Tehseen, M, Al-Amodi, A, Nguyen, P.Q, Savva, C.G, Hamdan, S.M, De Biasio, A. | | Deposit date: | 2024-06-20 | | Release date: | 2025-06-25 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structure of the human CTF18-RFC clamp loader bound to PCNA.
Biorxiv, 2025
|
|
9IN7
 
 | | True-atomic resolution crystal structure of the closed state of the viral channelrhodopsin OLPVR1 | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S)-2,3-dihydroxypropyl (9Z)-hexadec-9-enoate, EICOSANE, ... | | Authors: | Bukhdruker, S, Zabelskii, D, Astashkin, R, Gordeliy, V. | | Deposit date: | 2024-07-05 | | Release date: | 2025-03-12 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Ion-conducting and gating molecular mechanisms of channelrhodopsin revealed by true-atomic-resolution structures of open and closed states.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9IN8
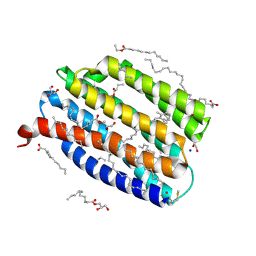 
 | | True-atomic resolution crystal structure of the open state of the viral channelrhodopsin OLPVR1 | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S)-2,3-dihydroxypropyl (9Z)-hexadec-9-enoate, EICOSANE, ... | | Authors: | Bukhdruker, S, Zabelskii, D, Astashkin, R, Bourenkov, G, Gordeliy, V. | | Deposit date: | 2024-07-05 | | Release date: | 2025-03-12 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Ion-conducting and gating molecular mechanisms of channelrhodopsin revealed by true-atomic-resolution structures of open and closed states.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9IN9
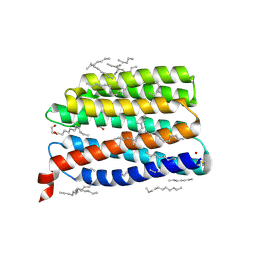 
 | | Structure of the viral channelrhodopsin OLPVR1 at low pH obtained by soaking | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S)-2,3-dihydroxypropyl (9Z)-hexadec-9-enoate, EICOSANE, ... | | Authors: | Bukhdruker, S, Zabelskii, D, Astashkin, R, Gordeliy, V. | | Deposit date: | 2024-07-05 | | Release date: | 2025-03-12 | | Last modified: | 2025-08-27 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Ion-conducting and gating molecular mechanisms of channelrhodopsin revealed by true-atomic-resolution structures of open and closed states.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9IUI
 
 | |
9IV9
 
 | | Cryo-EM structure of a truncated Nipah Virus L Protein bound by Phosphoprotein Tetramer | | Descriptor: | Phosphoprotein, RNA-directed RNA polymerase L, ZINC ION | | Authors: | Xue, L, Chang, T, Gui, J, Li, Z, Zhao, H, Zou, B, Li, M, He, J, Chen, X, Xiong, X. | | Deposit date: | 2024-07-23 | | Release date: | 2025-05-21 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (2.31 Å) | | Cite: | Cryo-EM structures of Nipah virus polymerase complex reveal highly varied interactions between L and P proteins among paramyxoviruses.
Protein Cell, 16, 2025
|
|
9IVA
 
 | | Cryo-EM structure of the full-length Nipah Virus L Protein bound by Phosphoprotein Tetramer | | Descriptor: | Phosphoprotein, RNA-directed RNA polymerase L, ZINC ION | | Authors: | Xue, L, Chang, T, Gui, J, Li, Z, Zhao, H, Zou, B, Li, M, He, J, Chen, X, Xiong, X. | | Deposit date: | 2024-07-23 | | Release date: | 2025-05-21 | | Last modified: | 2025-08-27 | | Method: | ELECTRON MICROSCOPY (2.52 Å) | | Cite: | Cryo-EM structures of Nipah virus polymerase complex reveal highly varied interactions between L and P proteins among paramyxoviruses.
Protein Cell, 16, 2025
|
|
9J5P
 
 | |
9J9H
 
 | |
9J9I
 
 | |
