1FVC
 
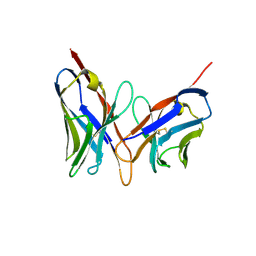 | | X-RAY STRUCTURES OF THE ANTIGEN-BINDING DOMAINS FROM THREE VARIANTS OF HUMANIZED ANTI-P185-HER2 ANTIBODY 4D5 AND COMPARISON WITH MOLECULAR MODELING | | Descriptor: | IGG1-KAPPA 4D5 FV (HEAVY CHAIN), IGG1-KAPPA 4D5 FV (LIGHT CHAIN) | | Authors: | Eigenbrot, C, Randal, M, Kossiakoff, A.A, Presta, L. | | Deposit date: | 1992-10-20 | | Release date: | 1993-10-31 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray structures of the antigen-binding domains from three variants of humanized anti-p185HER2 antibody 4D5 and comparison with molecular modeling.
J.Mol.Biol., 229, 1993
|
|
1FVD
 
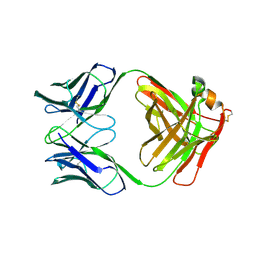 | | X-RAY STRUCTURES OF THE ANTIGEN-BINDING DOMAINS FROM THREE VARIANTS OF HUMANIZED ANTI-P185-HER2 ANTIBODY 4D5 AND COMPARISON WITH MOLECULAR MODELING | | Descriptor: | IGG1-KAPPA 4D5 FAB (HEAVY CHAIN), IGG1-KAPPA 4D5 FAB (LIGHT CHAIN) | | Authors: | Eigenbrot, C, Presta, L, Randal, M, Kossiakoff, A.A. | | Deposit date: | 1992-10-20 | | Release date: | 1993-10-31 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-ray structures of the antigen-binding domains from three variants of humanized anti-p185HER2 antibody 4D5 and comparison with molecular modeling.
J.Mol.Biol., 229, 1993
|
|
1FVE
 
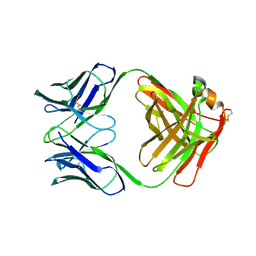 | | X-RAY STRUCTURES OF THE ANTIGEN-BINDING DOMAINS FROM THREE VARIANTS OF HUMANIZED ANTI-P185-HER2 ANTIBODY 4D5 AND COMPARISON WITH MOLECULAR MODELING | | Descriptor: | IGG1-KAPPA 4D5 FAB (HEAVY CHAIN), IGG1-KAPPA 4D5 FAB (LIGHT CHAIN) | | Authors: | Eigenbrot, C, Randal, M, Presta, L, Kossiakoff, A.A. | | Deposit date: | 1992-10-20 | | Release date: | 1993-10-31 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | X-ray structures of the antigen-binding domains from three variants of humanized anti-p185HER2 antibody 4D5 and comparison with molecular modeling.
J.Mol.Biol., 229, 1993
|
|
1FVF
 
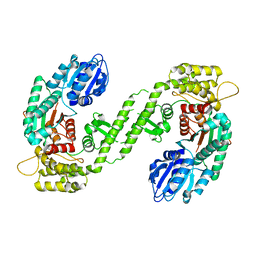 | |
1FVG
 
 | | CRYSTAL STRUCTURE OF BOVINE PEPTIDE METHIONINE SULFOXIDE REDUCTASE | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, PEPTIDE METHIONINE SULFOXIDE REDUCTASE | | Authors: | Lowther, W.T, Brot, N, Weissbach, H, Matthews, B.W. | | Deposit date: | 2000-09-19 | | Release date: | 2000-11-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure and mechanism of peptide methionine sulfoxide reductase, an "anti-oxidation" enzyme.
Biochemistry, 39, 2000
|
|
1FVH
 
 | |
1FVJ
 
 | |
1FVK
 
 | |
1FVO
 
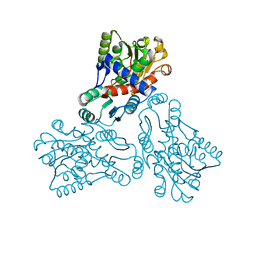 | | CRYSTAL STRUCTURE OF HUMAN ORNITHINE TRANSCARBAMYLASE COMPLEXED WITH CARBAMOYL PHOSPHATE | | Descriptor: | ORNITHINE TRANSCARBAMYLASE, PHOSPHORIC ACID MONO(FORMAMIDE)ESTER | | Authors: | Shi, D, Morizono, H, Yu, X, Allewell, N.M, Tuchman, M. | | Deposit date: | 2000-09-20 | | Release date: | 2001-04-04 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Human ornithine transcarbamylase: crystallographic insights into substrate recognition and conformational changes.
Biochem.J., 354, 2001
|
|
1FVR
 
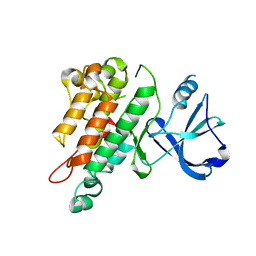 | | TIE2 KINASE DOMAIN | | Descriptor: | TYROSINE-PROTEIN KINASE TIE-2 | | Authors: | Shewchuk, L.M, Hassell, A.M, Ellis, B, Holmes, W.D, Davis, R, Horne, E.L, Kadwell, S.H, McKee, D.D, Moore, J.T. | | Deposit date: | 2000-09-20 | | Release date: | 2001-09-20 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of the Tie2 RTK domain: self-inhibition by the nucleotide binding loop, activation loop, and C-terminal tail.
Structure Fold.Des., 8, 2000
|
|
1FVT
 
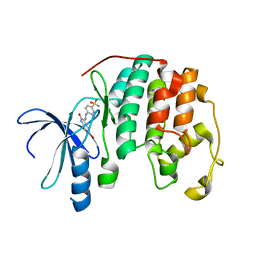 | | THE STRUCTURE OF CYCLIN-DEPENDENT KINASE 2 (CDK2) IN COMPLEX WITH AN OXINDOLE INHIBITOR | | Descriptor: | 4-[(2Z)-2-(5-bromo-2-oxo-1,2-dihydro-3H-indol-3-ylidene)hydrazinyl]benzene-1-sulfonamide, CELL DIVISION PROTEIN KINASE 2 | | Authors: | Davis, S.T, Benson, B.G, Bramson, H.N, Chapman, D.E, Dickerson, S.H, Dold, K.M, Eberwein, D.J, Edelstein, M, Frye, S.V, Gampe Jr, R.T, Griffin, R.J, Harris, P.A, Hassell, A.M, Holmes, W.D, Hunter, R.N, Knick, V.B, Lackey, K, Lovejoy, B, Luzzio, M.J, Murray, D, Parker, P, Rocque, W.J, Shewchuk, L, Veal, J.M, Walker, D.H, Kuyper, L.K. | | Deposit date: | 2000-09-20 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Prevention of chemotherapy-induced alopecia in rats by CDK inhibitors.
Science, 291, 2001
|
|
1FVV
 
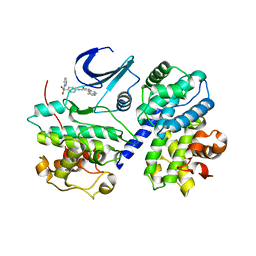 | | THE STRUCTURE OF CDK2/CYCLIN A IN COMPLEX WITH AN OXINDOLE INHIBITOR | | Descriptor: | 4-[(7-OXO-7H-THIAZOLO[5,4-E]INDOL-8-YLMETHYL)-AMINO]-N-PYRIDIN-2-YL-BENZENESULFONAMIDE, CYCLIN A, CYCLIN-DEPENDENT KINASE 2 | | Authors: | Davis, S.T, Benson, B.G, Bramson, H.N, Chapman, D.E, Dickerson, S.H, Dold, K.M, Eberwein, D.J, Edelstein, M, Frye, S.V, Gampe Jr, R.T, Griffin, R.J, Harris, P.A, Hassell, A.M, Holmes, W.D, Hunter, R.N, Knick, V.B, Lackey, K, Lovejoy, B, Luzzio, M.J, Murray, D, Parker, P, Rocque, W.J, Shewchuk, L, Veal, J.M, Walker, D.H, Kuyper, L.K. | | Deposit date: | 2000-09-20 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Prevention of chemotherapy-induced alopecia in rats by CDK inhibitors.
Science, 291, 2001
|
|
1FW1
 
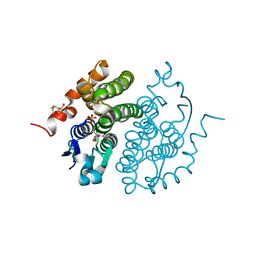 | | Glutathione transferase zeta/maleylacetoacetate isomerase | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, GLUTATHIONE, GLUTATHIONE TRANSFERASE ZETA, ... | | Authors: | Polekhina, G, Board, P.G, Blackburn, A.C, Parker, M.W. | | Deposit date: | 2000-09-20 | | Release date: | 2001-09-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of maleylacetoacetate isomerase/glutathione transferase zeta reveals the molecular basis for its remarkable catalytic promiscuity.
Biochemistry, 40, 2001
|
|
1FW4
 
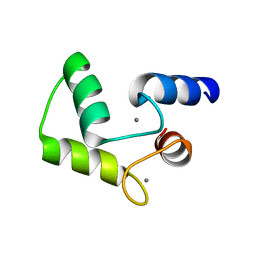 | |
1FW8
 
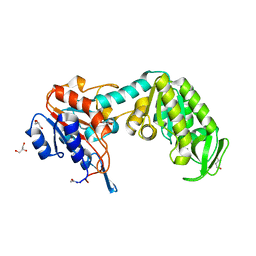 | | CIRCULARLY PERMUTED PHOSPHOGLYCERATE KINASE FROM YEAST: PGK P72 | | Descriptor: | GLYCEROL, Phosphoglycerate kinase | | Authors: | Tougard, P, Bizebard, T, Ritco-Vonsovici, M, Minard, P, Desmadril, M. | | Deposit date: | 2000-09-22 | | Release date: | 2001-03-22 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of a circularly permuted phosphoglycerate kinase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1FW9
 
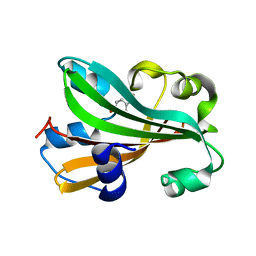 | | CHORISMATE LYASE WITH BOUND PRODUCT | | Descriptor: | CHORISMATE LYASE, P-HYDROXYBENZOIC ACID | | Authors: | Gallagher, D.T, Mayhew, M, Holden, M, Vilker, V, Howard, A. | | Deposit date: | 2000-09-22 | | Release date: | 2001-03-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The crystal structure of chorismate lyase shows a new fold and a tightly retained product.
Proteins, 44, 2001
|
|
1FWA
 
 | |
1FWB
 
 | |
1FWC
 
 | |
1FWD
 
 | |
1FWE
 
 | |
1FWF
 
 | |
1FWG
 
 | |
1FWH
 
 | |
1FWI
 
 | |
