7FJ1
 
 | | Cryo-EM structure of pseudorabies virus C-capsid | | Descriptor: | Capsid vertex component 1, DNA packaging tegument protein UL25, Major capsid protein, ... | | Authors: | Zheng, Q, Li, S, Zha, Z, Sun, H. | | Deposit date: | 2021-08-02 | | Release date: | 2022-06-22 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (4.43 Å) | | Cite: | Structures of pseudorabies virus capsids.
Nat Commun, 13, 2022
|
|
7FJ3
 
 | | Cryo-EM structure of PRV A-capid | | Descriptor: | Major capsid protein, Small capsomere-interacting protein, Triplex capsid protein 1, ... | | Authors: | Zheng, Q, Li, S, Zha, Z, Sun, H. | | Deposit date: | 2021-08-02 | | Release date: | 2022-06-22 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (4.53 Å) | | Cite: | Structures of pseudorabies virus capsids.
Nat Commun, 13, 2022
|
|
7FO8
 
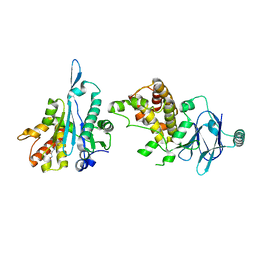 | | PanDDA analysis group deposition -- Aar2/RNaseH in complex with fragment P07H06 from the F2X-Universal Library | | Descriptor: | A1 cistron-splicing factor AAR2, N-[(1E)-2-(hydroxyamino)-2-oxoethylidene]benzamide, Pre-mRNA-splicing factor 8 | | Authors: | Barthel, T, Wollenhaupt, J, Lima, G.M.A, Wahl, M.C, Weiss, M.S. | | Deposit date: | 2022-08-26 | | Release date: | 2022-11-02 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Large-Scale Crystallographic Fragment Screening Expedites Compound Optimization and Identifies Putative Protein-Protein Interaction Sites.
J.Med.Chem., 65, 2022
|
|
7FTG
 
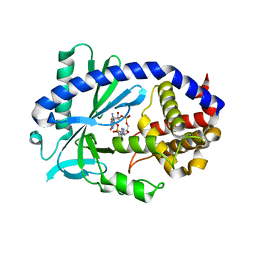 | |
7FW0
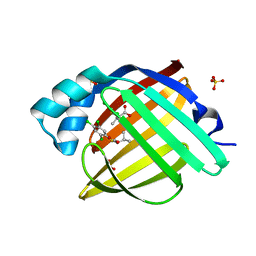 
 | | Crystal Structure of human FABP4 in complex with 3-methyl-2-(2,4,5-trichlorophenyl)sulfanylbutanoic acid, i.e. SMILES c1(c(cc(c(c1)Cl)S[C@@H](C(=O)O)C(C)C)Cl)Cl with IC50=3.5 microM | | Descriptor: | (2R)-3-methyl-2-[(2,4,5-trichlorophenyl)sulfanyl]butanoic acid, (2S)-3-methyl-2-[(2,4,5-trichlorophenyl)sulfanyl]butanoic acid, FORMIC ACID, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Richter, H, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2025-08-13 | | Method: | X-RAY DIFFRACTION (1.12 Å) | | Cite: | A high-resolution data set of fatty acid-binding protein structures. III. Unexpectedly high occurrence of wrong ligands.
Acta Crystallogr D Struct Biol, 81, 2025
|
|
7FYL
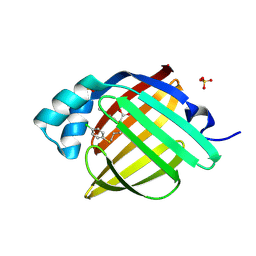 
 | | Crystal Structure of human FABP4 in complex with 6-(4-chlorophenyl)sulfanyl-2-(4-methylphenyl)sulfanylhexanoic acid, i.e. SMILES c1(S[C@@H](CCCCSc2ccc(cc2)Cl)C(=O)O)ccc(cc1)C with IC50=0.25724 microM | | Descriptor: | (2R)-6-[(4-chlorophenyl)sulfanyl]-2-[(4-methylphenyl)sulfanyl]hexanoic acid, (2S)-6-[(4-chlorophenyl)sulfanyl]-2-[(4-methylphenyl)sulfanyl]hexanoic acid, Fatty acid-binding protein, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Boehringer, M, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2025-08-13 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | A high-resolution data set of fatty acid-binding protein structures. III. Unexpectedly high occurrence of wrong ligands.
Acta Crystallogr D Struct Biol, 81, 2025
|
|
7FZF
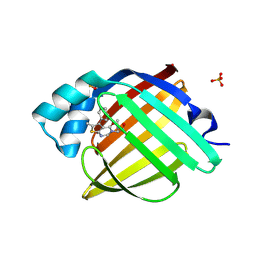 
 | | Crystal Structure of human FABP4 in complex with 2-(6-methyl-4-oxo-5-phenylthieno[2,3-d]pyrimidin-3-yl)butanoic acid, i.e. SMILES C12=C(N=CN(C1=O)[C@@H](C(=O)O)CC)SC(=C2c1ccccc1)C with IC50=7.52669 microM | | Descriptor: | (2R)-2-(6-methyl-4-oxo-5-phenylthieno[2,3-d]pyrimidin-3(4H)-yl)butanoic acid, (2S)-2-(6-methyl-4-oxo-5-phenylthieno[2,3-d]pyrimidin-3(4H)-yl)butanoic acid, Fatty acid-binding protein, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Brunner, M, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2025-08-13 | | Method: | X-RAY DIFFRACTION (1.09 Å) | | Cite: | A high-resolution data set of fatty acid-binding protein structures. III. Unexpectedly high occurrence of wrong ligands.
Acta Crystallogr D Struct Biol, 81, 2025
|
|
7GSA
 
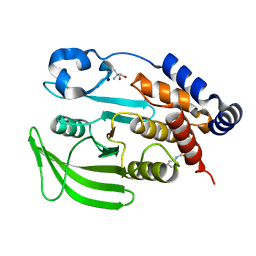 | |
7GSB
 
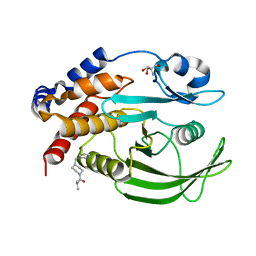 | |
7GSL
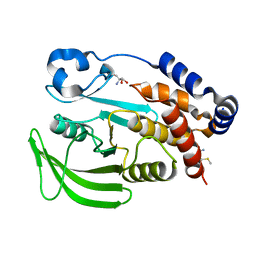 
 | |
7GSM
 
 | |
7GSO
 
 | |
7GSR
 
 | |
7GST
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOMB000056a | | Descriptor: | 1-(methanesulfonyl)-1,2,3,4-tetrahydroquinoline, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GSV
 
 | |
7GSW
 
 | |
7GSY
 
 | |
7GT0
 
 | |
7GT1
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOMB000209a | | Descriptor: | (2S)-2-(2-chloro-6-fluorophenyl)-2,3-dihydroquinazolin-4(1H)-one, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GT3
 
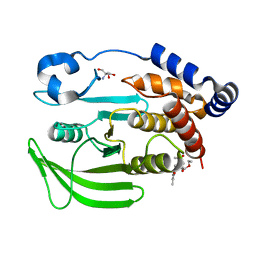 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000527a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1, ethyl (3R,3aS,8bS)-1-acetyl-5-methyl-2,3,3a,8b-tetrahydro-1H-[1]benzofuro[3,2-b]pyrrole-3-carboxylate | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GT4
 
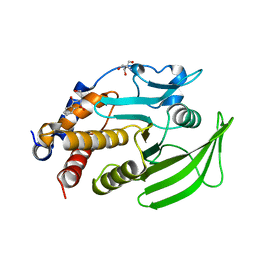 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000528a | | Descriptor: | (4R)-4-hydroxy-2-(2-hydroxyethyl)-3,4-dihydro-1lambda~6~,2-benzothiazine-1,1(2H)-dione, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GT5
 
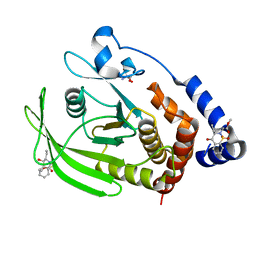 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000529a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1, methyl [(3R,4S)-3-ethyl-4-hydroxy-1,1-dioxo-3,4-dihydro-1lambda~6~,2-benzothiazin-2(1H)-yl]acetate | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GT6
 
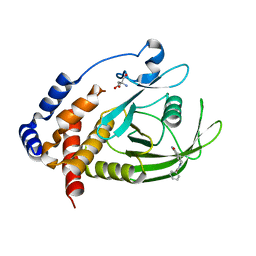 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000530a | | Descriptor: | (3aS,8aS)-6-benzoyloctahydropyrrolo[3,4-d]azepin-1(2H)-one, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GT7
 
 | |
7GT9
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMSOA000463b | | Descriptor: | (3R)-4-oxo-3,4-dihydro-2H-1-benzopyran-3-carbonitrile, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|







