+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4r44 | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Racemic crystal structure of a tetramolecular DNA G-quadruplex | ||||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / racemic DNA / racemates | Function / homology | : / DNA |  Function and homology information Function and homology informationBiological species | synthetic construct (others) | Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.695 Å MOLECULAR REPLACEMENT / Resolution: 2.695 Å  Authors AuthorsMandal, P.K. / Collie, G.W. / Kauffmann, B. / Huc, I. |  Citation Citation Journal: Angew.Chem.Int.Ed.Engl. / Year: 2014 Journal: Angew.Chem.Int.Ed.Engl. / Year: 2014Title: Racemic DNA crystallography. Authors: Mandal, P.K. / Collie, G.W. / Kauffmann, B. / Huc, I. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4r44.cif.gz 4r44.cif.gz | 58.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4r44.ent.gz pdb4r44.ent.gz | 43 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4r44.json.gz 4r44.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  4r44_validation.pdf.gz 4r44_validation.pdf.gz | 419.9 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  4r44_full_validation.pdf.gz 4r44_full_validation.pdf.gz | 420.8 KB | Display | |
| Data in XML |  4r44_validation.xml.gz 4r44_validation.xml.gz | 4.5 KB | Display | |
| Data in CIF |  4r44_validation.cif.gz 4r44_validation.cif.gz | 6.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/r4/4r44 https://data.pdbj.org/pub/pdb/validation_reports/r4/4r44 ftp://data.pdbj.org/pub/pdb/validation_reports/r4/4r44 ftp://data.pdbj.org/pub/pdb/validation_reports/r4/4r44 | HTTPS FTP |
-Related structure data
| Related structure data |  4r45C  4r47C  4r48C  4r49C  4r4aC  4r4dC  352dS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 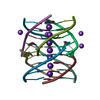
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 1880.251 Da / Num. of mol.: 16 / Source method: obtained synthetically / Details: Tetramolecular G-Quadruplex / Source: (synth.) synthetic construct (others) #2: Chemical | ChemComp-K / #3: Chemical | ChemComp-NA / #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.01 Å3/Da / Density % sol: 38.67 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 6.6 Details: 1 mM DNA, 50 mM potassium cacodylate, 40 mM potassium chloride, 100 mM magnesium chloride hexahydrate, 1.7 M 1,6-hexanediol, pH 6.6, VAPOR DIFFUSION, HANGING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å |
| Detector | Type: DECTRIS PILATUS 200K / Detector: PIXEL / Date: Dec 26, 2013 |
| Radiation | Monochromator: graphite / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.695→37.886 Å / Num. all: 12894 / Num. obs: 12597 / % possible obs: 97.69 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Redundancy: 1.7 % / Biso Wilson estimate: 49.77 Å2 / Rmerge(I) obs: 0.1643 / Net I/σ(I): 1.7 |
| Reflection shell | Resolution: 2.695→2.79 Å / Redundancy: 1.6 % / Rmerge(I) obs: 0.4338 / Mean I/σ(I) obs: 1.6 / Num. unique all: 1292 / % possible all: 97.34 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 352D Resolution: 2.695→37.886 Å / Cor.coef. Fo:Fc: 0.939 / Cor.coef. Fo:Fc free: 0.879 / SU B: 7.219 / SU ML: 0.153 / Cross valid method: THROUGHOUT / ESU R Free: 0.343 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN USED IF PRESENT IN THE INPUT
| |||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 27.827 Å2
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.695→37.886 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.695→2.765 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller









 PDBj
PDBj






