+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2jyv | ||||||
|---|---|---|---|---|---|---|---|
| Title | Human Granulin F | ||||||
 Components Components | Granulin-2 | ||||||
 Keywords Keywords |  CYTOKINE / Granulin C / epithelin / CYTOKINE / Granulin C / epithelin /  human / stack of hairpins / human / stack of hairpins /  Alternative splicing / Alternative splicing /  Glycoprotein / Polymorphism / Glycoprotein / Polymorphism /  Secreted Secreted | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of aspartic-type peptidase activity / positive regulation of inflammatory response to wounding / negative regulation of neutrophil activation / positive regulation of protein folding / microglial cell activation involved in immune response / positive regulation of lysosome organization / positive regulation of defense response to bacterium / negative regulation of microglial cell activation / lysosomal lumen acidification / astrocyte activation involved in immune response ...positive regulation of aspartic-type peptidase activity / positive regulation of inflammatory response to wounding / negative regulation of neutrophil activation / positive regulation of protein folding / microglial cell activation involved in immune response / positive regulation of lysosome organization / positive regulation of defense response to bacterium / negative regulation of microglial cell activation / lysosomal lumen acidification / astrocyte activation involved in immune response / lysosomal transport / lysosome organization / positive regulation of axon regeneration / negative regulation of respiratory burst involved in inflammatory response / positive regulation of endothelial cell migration /  cytokine activity / positive regulation of epithelial cell proliferation / cytokine activity / positive regulation of epithelial cell proliferation /  growth factor activity / growth factor activity /  trans-Golgi network / positive regulation of neuron apoptotic process / positive regulation of angiogenesis / azurophil granule lumen / late endosome / trans-Golgi network / positive regulation of neuron apoptotic process / positive regulation of angiogenesis / azurophil granule lumen / late endosome /  regulation of inflammatory response / protein-folding chaperone binding / negative regulation of neuron apoptotic process / regulation of inflammatory response / protein-folding chaperone binding / negative regulation of neuron apoptotic process /  lysosome / protein stabilization / lysosome / protein stabilization /  endosome / positive regulation of cell migration / lysosomal membrane / Neutrophil degranulation / endosome / positive regulation of cell migration / lysosomal membrane / Neutrophil degranulation /  Golgi apparatus / Golgi apparatus /  endoplasmic reticulum / endoplasmic reticulum /  signal transduction / signal transduction /  extracellular space / extracellular space /  RNA binding / extracellular exosome / extracellular region / RNA binding / extracellular exosome / extracellular region /  membrane / membrane /  plasma membrane plasma membraneSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Tolkatchev, D. / Wang, P. / Chen, Z. / Xu, P. / Ni, F. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2008 Journal: Protein Sci. / Year: 2008Title: Structure dissection of human progranulin identifies well-folded granulin/epithelin modules with unique functional activities. Authors: Tolkatchev, D. / Malik, S. / Vinogradova, A. / Wang, P. / Chen, Z. / Xu, P. / Bennett, H.P. / Bateman, A. / Ni, F. | ||||||
| History |
|
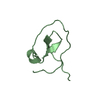
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2jyv.cif.gz 2jyv.cif.gz | 92.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2jyv.ent.gz pdb2jyv.ent.gz | 76.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2jyv.json.gz 2jyv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jy/2jyv https://data.pdbj.org/pub/pdb/validation_reports/jy/2jyv ftp://data.pdbj.org/pub/pdb/validation_reports/jy/2jyv ftp://data.pdbj.org/pub/pdb/validation_reports/jy/2jyv | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein |  / Granulin F / Granulin FMass: 7978.207 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: GRN / Plasmid: pET-32 / Production host: Homo sapiens (human) / Gene: GRN / Plasmid: pET-32 / Production host:   Escherichia coli (E. coli) / Strain (production host): Origami(DE3) / References: UniProt: P28799 Escherichia coli (E. coli) / Strain (production host): Origami(DE3) / References: UniProt: P28799 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||||||||||
| Sample conditions |
|
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 60 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller
















 PDBj
PDBj