+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1sen | ||||||
|---|---|---|---|---|---|---|---|
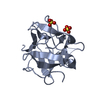
| Title | Endoplasmic reticulum protein Rp19 O95881 | ||||||
 Components Components | Thioredoxin-like protein p19 | ||||||
 Keywords Keywords | STRUCTURAL GENOMICS / UNKNOWN FUNCTION / Endoplasmic reticulum / Rp19 / PSI / Protein Structure Initiative / Southeast Collaboratory for Structural Genomics / SECSG | ||||||
| Function / homology |  Function and homology information Function and homology informationprotein-disulfide reductase (glutathione) / protein-disulfide reductase (glutathione) activity / protein-disulfide reductase activity / negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway / endoplasmic reticulum lumen / endoplasmic reticulum Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIR / Resolution: 1.199 Å MIR / Resolution: 1.199 Å | ||||||
 Authors Authors | Liu, Z.-J. / Chen, L. / Tempel, W. / Shah, A. / Lee, D. / Dailey, T.A. / Mayer, M.R. / Rose, J.P. / Richardson, D.C. / Richardson, J.S. ...Liu, Z.-J. / Chen, L. / Tempel, W. / Shah, A. / Lee, D. / Dailey, T.A. / Mayer, M.R. / Rose, J.P. / Richardson, D.C. / Richardson, J.S. / Dailey, H.A. / Wang, B.-C. / Southeast Collaboratory for Structural Genomics (SECSG) | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Endoplasmic reticulum protein Rp19 Authors: Liu, Z.-J. / Chen, L. / Tempel, W. / Shah, A. / Lee, D. / Rose, J.P. / Richardson, D.C. / Richardson, J.S. / Wang, B.-C. #1:  Journal: ACTA CRYSTALLOGR.,SECT.D / Year: 1997 Journal: ACTA CRYSTALLOGR.,SECT.D / Year: 1997Title: Refinement of Macromolecular Structures by the Maximum-Likelihood Method Authors: Murshudov, G.N. / Vagin, A.A. / Dodson, E.J. #2:  Journal: ACTA CRYSTALLOGR.,SECT.D / Year: 1999 Journal: ACTA CRYSTALLOGR.,SECT.D / Year: 1999Title: Automated MAD and MIR structure solution Authors: Terwilliger, T.C. / Berendzen, J. #3:  Journal: ACTA CRYSTALLOGR.,SECT.D / Year: 2000 Journal: ACTA CRYSTALLOGR.,SECT.D / Year: 2000Title: Maximum likelihood density modification Authors: Terwilliger, T.C. #4:  Journal: Nat.Struct.Biol. / Year: 1999 Journal: Nat.Struct.Biol. / Year: 1999Title: Automated protein model building combined with iterative structure refinement Authors: Perrakis, A. / Morris, R. / Lamzin, V.S. | ||||||
| History |
| ||||||
| Remark 300 | BIOMOLECULE THIS ENTRY CONTAINS THE CRYSTALLOGRAPHIC ASYMMETRIC UNIT WHICH CONSISTS OF 1 CHAIN(S). ...BIOMOLECULE THIS ENTRY CONTAINS THE CRYSTALLOGRAPHIC ASYMMETRIC UNIT WHICH CONSISTS OF 1 CHAIN(S). THE BIOLOGICAL UNIT IS UNKNOWN. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1sen.cif.gz 1sen.cif.gz | 40.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1sen.ent.gz pdb1sen.ent.gz | 29 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1sen.json.gz 1sen.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1sen_validation.pdf.gz 1sen_validation.pdf.gz | 420.3 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1sen_full_validation.pdf.gz 1sen_full_validation.pdf.gz | 420.6 KB | Display | |
| Data in XML |  1sen_validation.xml.gz 1sen_validation.xml.gz | 8.1 KB | Display | |
| Data in CIF |  1sen_validation.cif.gz 1sen_validation.cif.gz | 10.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/se/1sen https://data.pdbj.org/pub/pdb/validation_reports/se/1sen ftp://data.pdbj.org/pub/pdb/validation_reports/se/1sen ftp://data.pdbj.org/pub/pdb/validation_reports/se/1sen | HTTPS FTP |
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 18304.428 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) Homo sapiens (human)Description: The protein was cloned, expressed and purified by the SECSG human protein production group (T.A. Dailey, M. Mayer) under the direction of H.A. Dailey. Gene: TLP19 / Production host:  | ||||
|---|---|---|---|---|---|
| #2: Chemical | | #3: Chemical | #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.01 Å3/Da / Density % sol: 38.74 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: modified batch crystallization / pH: 7 Details: 12% PEG 4000, 100mM Na Hepes, 100mM NaCl, pH 7.0, modified batch crystallization, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1.07 Å / Beamline: 22-ID / Wavelength: 1.07 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Dec 19, 2003 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.07 Å / Relative weight: 1 |
| Reflection | Resolution: 1.199→38.633 Å / Num. all: 42206 / Num. obs: 42206 / Observed criterion σ(I): -3 |
-Phasing
| Phasing | Method:  MIR MIR | |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phasing dm | FOM : 0.53 / FOM acentric: 0.51 / FOM centric: 0.6 / Reflection: 7438 / Reflection acentric: 6336 / Reflection centric: 1102 | |||||||||||||||||||||||||||||||||||||||||||||||||
| Phasing dm shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||
| Phasing MIR | Resolution: 2.5→20 Å / FOM: 0.33 / Reflection: 5250 | |||||||||||||||||||||||||||||||||||||||||||||||||
| Phasing MIR shell |
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR / Resolution: 1.199→38.633 Å / Cor.coef. Fo:Fc: 0.965 / SU R Cruickshank DPI: 0.041 / Cross valid method: THROUGHOUT / Stereochemistry target values: Engh & Huber MIR / Resolution: 1.199→38.633 Å / Cor.coef. Fo:Fc: 0.965 / SU R Cruickshank DPI: 0.041 / Cross valid method: THROUGHOUT / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: BABINET MODEL PLUS MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 12.564 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.199→38.633 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller












 PDBj
PDBj







