+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1kt3 | ||||||
|---|---|---|---|---|---|---|---|
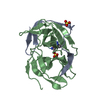
| Title | Crystal structure of bovine holo-RBP at pH 2.0 | ||||||
 Components Components | Plasma retinol-binding protein | ||||||
 Keywords Keywords | TRANSPORT PROTEIN / RBP / retinol binding | ||||||
| Function / homology |  Function and homology information Function and homology informationretinol transport / retinol transmembrane transporter activity / retinal binding / retinol binding / extracellular space Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.4 Å MOLECULAR REPLACEMENT / Resolution: 1.4 Å | ||||||
 Authors Authors | Calderone, V. / Berni, R. / Zanotti, G. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2003 Journal: J.Mol.Biol. / Year: 2003Title: High-resolution Structures of Retinol-binding Protein in Complex with Retinol: pH-induced Protein Structural Changes in the Crystal State Authors: Calderone, V. / Berni, R. / Zanotti, G. #1:  Journal: Proteins / Year: 1990 Journal: Proteins / Year: 1990Title: Crystallographic Refinement of Human Serum Retinol Binding Protein at 2A Resolution. Authors: Cowan, S.W. / Newcomer, M.E. / Jones, T.A. #2:  Journal: J.Biol.Chem. / Year: 1993 Journal: J.Biol.Chem. / Year: 1993Title: Crystal Structure of Liganded and Unliganded Forms of Bovine Plasma Retinol-Binding Protein Authors: Zanotti, G. / Berni, R. / Monaco, H.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1kt3.cif.gz 1kt3.cif.gz | 95 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1kt3.ent.gz pdb1kt3.ent.gz | 71.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1kt3.json.gz 1kt3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1kt3_validation.pdf.gz 1kt3_validation.pdf.gz | 434.4 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1kt3_full_validation.pdf.gz 1kt3_full_validation.pdf.gz | 440.4 KB | Display | |
| Data in XML |  1kt3_validation.xml.gz 1kt3_validation.xml.gz | 6.5 KB | Display | |
| Data in CIF |  1kt3_validation.cif.gz 1kt3_validation.cif.gz | 9.8 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kt/1kt3 https://data.pdbj.org/pub/pdb/validation_reports/kt/1kt3 ftp://data.pdbj.org/pub/pdb/validation_reports/kt/1kt3 ftp://data.pdbj.org/pub/pdb/validation_reports/kt/1kt3 | HTTPS FTP |
-Related structure data
| Related structure data |  1kt4C  1kt5C  1kt6C  1kt7C  1hbpS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 21095.654 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  |
|---|---|
| #2: Chemical | ChemComp-RTL / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.94 Å3/Da / Density % sol: 43.23 % | |||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 2 Details: 50 mM Na citrate, 100 mM NaCl, pH 2.0, VAPOR DIFFUSION, SITTING DROP, temperature 293K | |||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 7 / Method: vapor diffusion / Details: used microseeding | |||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-4 / Wavelength: 0.9393 Å / Beamline: ID14-4 / Wavelength: 0.9393 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Feb 10, 2001 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9393 Å / Relative weight: 1 |
| Reflection | Resolution: 1.399→34.246 Å / Num. all: 32419 / Num. obs: 32419 / % possible obs: 87.8 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 3.6 % / Rmerge(I) obs: 0.069 / Rsym value: 0.069 / Net I/σ(I): 5.7 |
| Reflection shell | Resolution: 1.4→1.47 Å / Redundancy: 2.3 % / Rmerge(I) obs: 0.464 / Mean I/σ(I) obs: 1 / Num. unique all: 1249 / Rsym value: 0.464 / % possible all: 42.7 |
| Reflection | *PLUS Highest resolution: 1.39 Å / Lowest resolution: 34.3 Å / Num. measured all: 334447 |
| Reflection shell | *PLUS Highest resolution: 1.39 Å / % possible obs: 42.7 % / Rmerge(I) obs: 0.382 / Mean I/σ(I) obs: 1.1 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1HBP Resolution: 1.4→34.246 Å / Num. parameters: 14484 / Num. restraintsaints: 17663 / Cross valid method: FREE R / σ(F): 0 / Stereochemistry target values: ENGH AND HUBER
| |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 0 / Occupancy sum hydrogen: 0 / Occupancy sum non hydrogen: 1609 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.4→34.246 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL / Version: 97 / Classification: refinement | |||||||||||||||||||||||||||||||||
| Refinement | *PLUS Num. reflection obs: 26941 / Num. reflection Rfree: 1347 / % reflection Rfree: 5 % / Rfactor Rfree: 0.2435 / Rfactor Rwork: 0.1421 | |||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller














 PDBj
PDBj




