+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1gjh | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
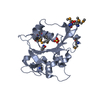
| Title | HUMAN BCL-2, ISOFORM 2 | |||||||||
 Components Components | PROTEIN (APOPTOSIS REGULATOR BCL-2 WITH PUTATIVE FLEXIBLE LOOP REPLACED WITH A PORTION OF APOPTOSIS REGULATOR BCL-X PROTEIN) | |||||||||
 Keywords Keywords |  APOPTOSIS APOPTOSIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of cellular pH reduction / negative regulation of retinal cell programmed cell death / pigment granule organization / channel inhibitor activity / CD8-positive, alpha-beta T cell lineage commitment / BAD-BCL-2 complex / regulation of glycoprotein biosynthetic process / melanin metabolic process / positive regulation of skeletal muscle fiber development / positive regulation of melanocyte differentiation ...negative regulation of cellular pH reduction / negative regulation of retinal cell programmed cell death / pigment granule organization / channel inhibitor activity / CD8-positive, alpha-beta T cell lineage commitment / BAD-BCL-2 complex / regulation of glycoprotein biosynthetic process / melanin metabolic process / positive regulation of skeletal muscle fiber development / positive regulation of melanocyte differentiation / myeloid cell apoptotic process / osteoblast proliferation / cochlear nucleus development / mesenchymal cell development / retinal cell programmed cell death / positive regulation of neuron maturation / negative regulation of osteoblast proliferation / gland morphogenesis /  renal system process / apoptotic process in bone marrow cell / T cell apoptotic process / regulation of cell-matrix adhesion / stem cell development / negative regulation of calcium ion transport into cytosol / melanocyte differentiation / ear development / The NLRP1 inflammasome / SARS-CoV-1-mediated effects on programmed cell death / dendritic cell apoptotic process / lymphoid progenitor cell differentiation / dendritic cell proliferation / positive regulation of mononuclear cell proliferation / negative regulation of myeloid cell apoptotic process / negative regulation of epithelial cell apoptotic process / regulation of nitrogen utilization / BH3-only proteins associate with and inactivate anti-apoptotic BCL-2 members / B cell apoptotic process / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage / glomerulus development / negative regulation of T cell apoptotic process / negative regulation of dendritic cell apoptotic process / oocyte development / neuron maturation / positive regulation of multicellular organism growth / negative regulation of execution phase of apoptosis / metanephros development / negative regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway / regulation of viral genome replication / renal system process / apoptotic process in bone marrow cell / T cell apoptotic process / regulation of cell-matrix adhesion / stem cell development / negative regulation of calcium ion transport into cytosol / melanocyte differentiation / ear development / The NLRP1 inflammasome / SARS-CoV-1-mediated effects on programmed cell death / dendritic cell apoptotic process / lymphoid progenitor cell differentiation / dendritic cell proliferation / positive regulation of mononuclear cell proliferation / negative regulation of myeloid cell apoptotic process / negative regulation of epithelial cell apoptotic process / regulation of nitrogen utilization / BH3-only proteins associate with and inactivate anti-apoptotic BCL-2 members / B cell apoptotic process / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage / glomerulus development / negative regulation of T cell apoptotic process / negative regulation of dendritic cell apoptotic process / oocyte development / neuron maturation / positive regulation of multicellular organism growth / negative regulation of execution phase of apoptosis / metanephros development / negative regulation of mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway / regulation of viral genome replication /  focal adhesion assembly / negative regulation of motor neuron apoptotic process / endoplasmic reticulum calcium ion homeostasis / negative regulation of ossification / focal adhesion assembly / negative regulation of motor neuron apoptotic process / endoplasmic reticulum calcium ion homeostasis / negative regulation of ossification /  fertilization / negative regulation of B cell apoptotic process / response to iron ion / negative regulation of protein localization to plasma membrane / response to UV-B / negative regulation of mitochondrial depolarization / regulation of mitochondrial membrane permeability / regulation of growth / calcium ion transport into cytosol / channel activity / motor neuron apoptotic process / axon regeneration / fertilization / negative regulation of B cell apoptotic process / response to iron ion / negative regulation of protein localization to plasma membrane / response to UV-B / negative regulation of mitochondrial depolarization / regulation of mitochondrial membrane permeability / regulation of growth / calcium ion transport into cytosol / channel activity / motor neuron apoptotic process / axon regeneration /  Bcl-2 family protein complex / epithelial cell apoptotic process / smooth muscle cell migration / intrinsic apoptotic signaling pathway in response to oxidative stress / organ growth / NFE2L2 regulating tumorigenic genes / digestive tract morphogenesis / response to cycloheximide / branching involved in ureteric bud morphogenesis / cellular response to organic substance / hair follicle morphogenesis / negative regulation of G1/S transition of mitotic cell cycle / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / cellular response to alkaloid / B cell lineage commitment / STAT5 activation downstream of FLT3 ITD mutants / hepatocyte apoptotic process / positive regulation of smooth muscle cell migration / B cell proliferation / pore complex / negative regulation of reproductive process / negative regulation of developmental process / negative regulation of release of cytochrome c from mitochondria / T cell homeostasis / Bcl-2 family protein complex / epithelial cell apoptotic process / smooth muscle cell migration / intrinsic apoptotic signaling pathway in response to oxidative stress / organ growth / NFE2L2 regulating tumorigenic genes / digestive tract morphogenesis / response to cycloheximide / branching involved in ureteric bud morphogenesis / cellular response to organic substance / hair follicle morphogenesis / negative regulation of G1/S transition of mitotic cell cycle / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / cellular response to alkaloid / B cell lineage commitment / STAT5 activation downstream of FLT3 ITD mutants / hepatocyte apoptotic process / positive regulation of smooth muscle cell migration / B cell proliferation / pore complex / negative regulation of reproductive process / negative regulation of developmental process / negative regulation of release of cytochrome c from mitochondria / T cell homeostasis /  BH3 domain binding / apoptotic mitochondrial changes / germ cell development / regulation of calcium ion transport / negative regulation of apoptotic signaling pathway / B cell homeostasis / BH3 domain binding / apoptotic mitochondrial changes / germ cell development / regulation of calcium ion transport / negative regulation of apoptotic signaling pathway / B cell homeostasis /  humoral immune response / negative regulation of anoikis / extrinsic apoptotic signaling pathway via death domain receptors / negative regulation of extrinsic apoptotic signaling pathway in absence of ligand / intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress / Estrogen-dependent nuclear events downstream of ESR-membrane signaling humoral immune response / negative regulation of anoikis / extrinsic apoptotic signaling pathway via death domain receptors / negative regulation of extrinsic apoptotic signaling pathway in absence of ligand / intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress / Estrogen-dependent nuclear events downstream of ESR-membrane signalingSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | |||||||||
 Authors Authors | Petros, A.M. / Medek, A. / Nettesheim, D.G. / Kim, D.H. / Yoon, H.S. / Swift, K. / Matayoshi, E.D. / Oltersdorf, T. / Fesik, S.W. | |||||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2001 Journal: Proc.Natl.Acad.Sci.USA / Year: 2001Title: Solution structure of the antiapoptotic protein bcl-2. Authors: Petros, A.M. / Medek, A. / Nettesheim, D.G. / Kim, D.H. / Yoon, H.S. / Swift, K. / Matayoshi, E.D. / Oltersdorf, T. / Fesik, S.W. #1:  Journal: Cell(Cambridge,Mass.) / Year: 1986 Journal: Cell(Cambridge,Mass.) / Year: 1986Title: Cloning and Structural Analysis of Cdnas for Bcl-2 and a Hybrid Bcl-2/Immunoglobulin Transcript Resulting from the T(14;18) Translocation Authors: Cleary, M.L. / Smith, S.D. / Sklar, J. #2:  Journal: Embo J. / Year: 1988 Journal: Embo J. / Year: 1988Title: Alternative Promoters and Exons, Somatic Mutation and Deregulation of the Bcl-2-Ig Fusion Gene in Lymphoma Authors: Seto, M. / Jaeger, U. / Hockett, R.D. / Grainger, W. / Bennett, S. / Goldman, P. / Korsmeyer, S.J. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1gjh.cif.gz 1gjh.cif.gz | 68.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1gjh.ent.gz pdb1gjh.ent.gz | 51.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1gjh.json.gz 1gjh.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gj/1gjh https://data.pdbj.org/pub/pdb/validation_reports/gj/1gjh ftp://data.pdbj.org/pub/pdb/validation_reports/gj/1gjh ftp://data.pdbj.org/pub/pdb/validation_reports/gj/1gjh | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 19357.557 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) / References: UniProt: P10415, UniProt: Q07817 Escherichia coli (E. coli) / References: UniProt: P10415, UniProt: Q07817 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 20 mM / pH: 7.80 / Pressure: AMBIENT / Temperature: 298.00 K | |||||||||
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software | Name: CNX / Version: 2000 / Developer: BRUNGER / Classification: refinement |
|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 |
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller















 PDBj
PDBj




