[English] 日本語
 Yorodumi
Yorodumi- PDB-1f8z: NMR STRUCTURE OF THE SIXTH LIGAND-BINDING MODULE OF THE LDL RECEPTOR -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1f8z | ||||||
|---|---|---|---|---|---|---|---|
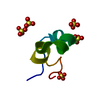
| Title | NMR STRUCTURE OF THE SIXTH LIGAND-BINDING MODULE OF THE LDL RECEPTOR | ||||||
 Components Components | LOW-DENSITY LIPOPROTEIN RECEPTOR LDL receptor LDL receptor | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN /  LDL receptor / ligand-binding domain / calcium-binding / LDL receptor / ligand-binding domain / calcium-binding /  familial hypercholesterolemia familial hypercholesterolemia | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of phosphatidylcholine catabolic process / plasma lipoprotein particle clearance / positive regulation of lysosomal protein catabolic process / very-low-density lipoprotein particle receptor activity / negative regulation of astrocyte activation / receptor-mediated endocytosis involved in cholesterol transport / response to caloric restriction / negative regulation of microglial cell activation / Chylomicron clearance / PCSK9-LDLR complex ...regulation of phosphatidylcholine catabolic process / plasma lipoprotein particle clearance / positive regulation of lysosomal protein catabolic process / very-low-density lipoprotein particle receptor activity / negative regulation of astrocyte activation / receptor-mediated endocytosis involved in cholesterol transport / response to caloric restriction / negative regulation of microglial cell activation / Chylomicron clearance / PCSK9-LDLR complex / cholesterol import / negative regulation of receptor recycling / low-density lipoprotein particle clearance / clathrin heavy chain binding / low-density lipoprotein particle receptor activity / high-density lipoprotein particle clearance / lipoprotein catabolic process / intestinal cholesterol absorption / positive regulation of triglyceride biosynthetic process / negative regulation of low-density lipoprotein particle clearance / regulation of protein metabolic process / low-density lipoprotein particle / low-density lipoprotein particle binding / amyloid-beta clearance by cellular catabolic process / LDL clearance / negative regulation of amyloid fibril formation / phospholipid transport / cholesterol transport / endolysosome membrane / negative regulation of protein metabolic process / artery morphogenesis / regulation of cholesterol metabolic process / cellular response to fatty acid /  lipoprotein particle binding / amyloid-beta clearance / sorting endosome / cellular response to low-density lipoprotein particle stimulus / lipoprotein particle binding / amyloid-beta clearance / sorting endosome / cellular response to low-density lipoprotein particle stimulus /  long-term memory / Retinoid metabolism and transport / long-term memory / Retinoid metabolism and transport /  phagocytosis / phagocytosis /  clathrin-coated pit / somatodendritic compartment / cholesterol metabolic process / clathrin-coated pit / somatodendritic compartment / cholesterol metabolic process /  receptor-mediated endocytosis / cholesterol homeostasis / clathrin-coated endocytic vesicle membrane / lipid metabolic process / positive regulation of inflammatory response / receptor-mediated endocytosis / cholesterol homeostasis / clathrin-coated endocytic vesicle membrane / lipid metabolic process / positive regulation of inflammatory response /  endocytosis / late endosome / Cargo recognition for clathrin-mediated endocytosis / virus receptor activity / apical part of cell / endocytosis / late endosome / Cargo recognition for clathrin-mediated endocytosis / virus receptor activity / apical part of cell /  Clathrin-mediated endocytosis / Clathrin-mediated endocytosis /  amyloid-beta binding / basolateral plasma membrane / amyloid-beta binding / basolateral plasma membrane /  protease binding / protease binding /  lysosome / molecular adaptor activity / lysosome / molecular adaptor activity /  receptor complex / receptor complex /  early endosome / endosome membrane / external side of plasma membrane / negative regulation of gene expression / early endosome / endosome membrane / external side of plasma membrane / negative regulation of gene expression /  calcium ion binding / positive regulation of gene expression / calcium ion binding / positive regulation of gene expression /  Golgi apparatus / Golgi apparatus /  cell surface / cell surface /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | ||||||
| Method |  SOLUTION NMR / Structure determination by Torsion angle dynamics, SOLUTION NMR / Structure determination by Torsion angle dynamics,  Molecular dynamics Molecular dynamics | ||||||
 Authors Authors | Clayton, D.J. / Brereton, I.M. / Kroon, P.A. / Smith, R. | ||||||
 Citation Citation |  Journal: FEBS Lett. / Year: 2000 Journal: FEBS Lett. / Year: 2000Title: Three-dimensional NMR structure of the sixth ligand-binding module of the human LDL receptor: comparison of two adjacent modules with different ligand binding specificities. Authors: Clayton, D. / Brereton, I.M. / Kroon, P.A. / Smith, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1f8z.cif.gz 1f8z.cif.gz | 220.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1f8z.ent.gz pdb1f8z.ent.gz | 189.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1f8z.json.gz 1f8z.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/f8/1f8z https://data.pdbj.org/pub/pdb/validation_reports/f8/1f8z ftp://data.pdbj.org/pub/pdb/validation_reports/f8/1f8z ftp://data.pdbj.org/pub/pdb/validation_reports/f8/1f8z | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide |  LDL receptor LDL receptorMass: 4392.759 Da / Num. of mol.: 1 / Fragment: SIXTH LIGAND-BINDING MODULE / Source method: obtained synthetically / Details: Peptide synthesis - based on the human sequence / References: UniProt: P01130 |
|---|---|
| #2: Chemical | ChemComp-CA / |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 10 mM / pH: 5.50 / Pressure: 1 atm / Temperature: 298 K | |||||||||
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker DMX / Manufacturer: Bruker / Model : DMX / Field strength: 750 MHz : DMX / Field strength: 750 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: Structure determination by Torsion angle dynamics,  Molecular dynamics Molecular dynamicsSoftware ordinal: 1 Details: Structures are based on 552 distance constraints, 18 backbone angle restraints and 14 sidechain angle restraints | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations, structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller











 PDBj
PDBj













