+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-3561 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
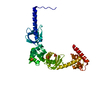
| タイトル | map of the RNA polymerase lambda-based antitermination complex solved by cryo-EM | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報bacterial-type RNA polymerase core enzyme binding / RNA stem-loop binding /  RNA polymerase binding / transcription elongation-coupled chromatin remodeling / RNA polymerase binding / transcription elongation-coupled chromatin remodeling /  RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly ...bacterial-type RNA polymerase core enzyme binding / RNA stem-loop binding / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly ...bacterial-type RNA polymerase core enzyme binding / RNA stem-loop binding /  RNA polymerase binding / transcription elongation-coupled chromatin remodeling / RNA polymerase binding / transcription elongation-coupled chromatin remodeling /  RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / transcription antitermination factor activity, RNA binding / bacterial-type flagellum-dependent cell motility / nitrate assimilation / transcription elongation factor complex / regulation of DNA-templated transcription elongation / transcription antitermination / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / transcription antitermination factor activity, RNA binding / bacterial-type flagellum-dependent cell motility / nitrate assimilation / transcription elongation factor complex / regulation of DNA-templated transcription elongation / transcription antitermination /  運動性 / DNA-templated transcription termination / DNA-templated transcription initiation / 運動性 / DNA-templated transcription termination / DNA-templated transcription initiation /  ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity /  ポリメラーゼ / cytosolic small ribosomal subunit / ポリメラーゼ / cytosolic small ribosomal subunit /  リボソーム生合成 / protein complex oligomerization / cytoplasmic translation / small ribosomal subunit / response to heat / protein-containing complex assembly / intracellular iron ion homeostasis / リボソーム生合成 / protein complex oligomerization / cytoplasmic translation / small ribosomal subunit / response to heat / protein-containing complex assembly / intracellular iron ion homeostasis /  tRNA binding / tRNA binding /  single-stranded RNA binding / single-stranded RNA binding /  protein dimerization activity / structural constituent of ribosome / DNA-binding transcription factor activity / protein domain specific binding / protein dimerization activity / structural constituent of ribosome / DNA-binding transcription factor activity / protein domain specific binding /  nucleotide binding / response to antibiotic / regulation of transcription by RNA polymerase II / magnesium ion binding / nucleotide binding / response to antibiotic / regulation of transcription by RNA polymerase II / magnesium ion binding /  DNA binding / DNA binding /  RNA binding / zinc ion binding / RNA binding / zinc ion binding /  生体膜 / 生体膜 /  細胞質基質 / 細胞質基質 /  細胞質 細胞質類似検索 - 分子機能 | |||||||||
| 生物種 |   Escherichia coli (大腸菌) / Escherichia coli (大腸菌) /   Escherichia phage lambda (λファージ) / Escherichia phage lambda (λファージ) /   Escherichia coli K-12 (大腸菌) Escherichia coli K-12 (大腸菌) | |||||||||
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 9.8 Å クライオ電子顕微鏡法 / 解像度: 9.8 Å | |||||||||
 データ登録者 データ登録者 | Said N / Krupp F | |||||||||
 引用 引用 |  ジャーナル: Nat Microbiol / 年: 2017 ジャーナル: Nat Microbiol / 年: 2017タイトル: Structural basis for λN-dependent processive transcription antitermination. 著者: Nelly Said / Ferdinand Krupp / Ekaterina Anedchenko / Karine F Santos / Olexandr Dybkov / Yong-Heng Huang / Chung-Tien Lee / Bernhard Loll / Elmar Behrmann / Jörg Bürger / Thorsten Mielke / ...著者: Nelly Said / Ferdinand Krupp / Ekaterina Anedchenko / Karine F Santos / Olexandr Dybkov / Yong-Heng Huang / Chung-Tien Lee / Bernhard Loll / Elmar Behrmann / Jörg Bürger / Thorsten Mielke / Justus Loerke / Henning Urlaub / Christian M T Spahn / Gert Weber / Markus C Wahl /  要旨: λN-mediated processive antitermination constitutes a paradigmatic transcription regulatory event, during which phage protein λN, host factors NusA, NusB, NusE and NusG, and an RNA nut site render ...λN-mediated processive antitermination constitutes a paradigmatic transcription regulatory event, during which phage protein λN, host factors NusA, NusB, NusE and NusG, and an RNA nut site render elongating RNA polymerase termination-resistant. The structural basis of the process has so far remained elusive. Here we describe a crystal structure of a λN-NusA-NusB-NusE-nut site complex and an electron cryo-microscopic structure of a complete transcription antitermination complex, comprising RNA polymerase, DNA, nut site RNA, all Nus factors and λN, validated by crosslinking/mass spectrometry. Due to intrinsic disorder, λN can act as a multiprotein/RNA interaction hub, which, together with nut site RNA, arranges NusA, NusB and NusE into a triangular complex. This complex docks via the NusA N-terminal domain and the λN C-terminus next to the RNA exit channel on RNA polymerase. Based on the structures, comparative crosslinking analyses and structure-guided mutagenesis, we hypothesize that λN mounts a multipronged strategy to reprogram the transcriptional machinery, which may include (1) the λN C terminus clamping the RNA exit channel, thus stabilizing the DNA:RNA hybrid; (2) repositioning of NusA and RNAP elements, thus redirecting nascent RNA and sequestering the upstream branch of a terminator hairpin; and (3) hindering RNA engagement of termination factor ρ and/or obstructing ρ translocation on the transcript. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_3561.map.gz emd_3561.map.gz | 48.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-3561-v30.xml emd-3561-v30.xml emd-3561.xml emd-3561.xml | 25.4 KB 25.4 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_3561.png emd_3561.png | 137.3 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3561 http://ftp.pdbj.org/pub/emdb/structures/EMD-3561 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3561 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3561 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_3561.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_3561.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.28 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : lambdaN-dependent antitermination complex
+超分子 #1: lambdaN-dependent antitermination complex
+分子 #1: Antitermination protein N
+分子 #3: DNA-directed RNA polymerase subunit alpha
+分子 #4: DNA-directed RNA polymerase subunit beta
+分子 #5: DNA-directed RNA polymerase subunit beta'
+分子 #6: 30S ribosomal protein S10
+分子 #7: Transcription termination/antitermination protein NusG
+分子 #11: Transcription antitermination protein NusB
+分子 #12: Transcription termination/antitermination protein NusA
+分子 #13: DNA-directed RNA polymerase subunit omega
+分子 #2: nascent RNA
+分子 #8: RNA transcription bubble
+分子 #9: DNAI
+分子 #10: DNAII
+分子 #14: MAGNESIUM ION
+分子 #15: ZINC ION
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
| 詳細 | 10 mM HEPES-NaOH, pH 7.5, 50 mM NaCl, 1 mM DTT |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI POLARA 300 |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: SUPER-RESOLUTION / 平均電子線量: 1.0 e/Å2 |
| 実験機器 |  モデル: Tecnai Polara / 画像提供: FEI Company |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: INSILICO MODEL 詳細: Generate from class averages using the VIPER algorithm. |
|---|---|
| 初期 角度割当 | タイプ: PROJECTION MATCHING |
| 最終 角度割当 | タイプ: PROJECTION MATCHING |
| 最終 再構成 | 想定した対称性 - 点群: C1 (非対称) / 解像度のタイプ: BY AUTHOR / 解像度: 9.8 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 使用した粒子像数: 23983 |
 ムービー
ムービー コントローラー
コントローラー