登録情報 データベース : EMDB / ID : EMD-0694タイトル Cryo-EM structure of human MLL1-NCP complex, binding mode1 Main map of human MLL1-NCP complex, binding mode1 複合体 : Human MLL1 complex associated with an unmodified nucleosome, binding mode 1複合体 : Human MLL1 complex複合体 : unmodified nucleosomeタンパク質・ペプチド : x 4種DNA : x 2種リガンド : x 4種 / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
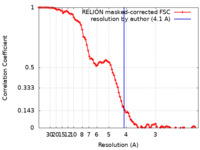
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト) / Xenopus laevis (アフリカツメガエル) / synthetic construct (人工物) 手法 / / 解像度 : 4.1 Å Huang J / Xue H 資金援助 Organization Grant number 国 National Science Foundation (China)
ジャーナル : Nature / 年 : 2019タイトル : Structural basis of nucleosome recognition and modification by MLL methyltransferases.著者 : Han Xue / Tonghui Yao / Mi Cao / Guanjun Zhu / Yan Li / Guiyong Yuan / Yong Chen / Ming Lei / Jing Huang / 要旨 : Methyltransferases of the mixed-lineage leukaemia (MLL) family-which include MLL1, MLL2, MLL3, MLL4, SET1A and SET1B-implement methylation of histone H3 on lysine 4 (H3K4), and have critical and ... Methyltransferases of the mixed-lineage leukaemia (MLL) family-which include MLL1, MLL2, MLL3, MLL4, SET1A and SET1B-implement methylation of histone H3 on lysine 4 (H3K4), and have critical and distinct roles in the regulation of transcription in haematopoiesis, adipogenesis and development. The C-terminal catalytic SET (Su(var.)3-9, enhancer of zeste and trithorax) domains of MLL proteins are associated with a common set of regulatory factors (WDR5, RBBP5, ASH2L and DPY30) to achieve specific activities. Current knowledge of the regulation of MLL activity is limited to the catalysis of histone H3 peptides, and how H3K4 methyl marks are deposited on nucleosomes is poorly understood. H3K4 methylation is stimulated by mono-ubiquitination of histone H2B on lysine 120 (H2BK120ub1), a prevalent histone H2B mark that disrupts chromatin compaction and favours open chromatin structures, but the underlying mechanism remains unknown. Here we report cryo-electron microscopy structures of human MLL1 and MLL3 catalytic modules associated with nucleosome core particles that contain H2BK120ub1 or unmodified H2BK120. These structures demonstrate that the MLL1 and MLL3 complexes both make extensive contacts with the histone-fold and DNA regions of the nucleosome; this allows ease of access to the histone H3 tail, which is essential for the efficient methylation of H3K4. The H2B-conjugated ubiquitin binds directly to RBBP5, orienting the association between MLL1 or MLL3 and the nucleosome. The MLL1 and MLL3 complexes display different structural organizations at the interface between the WDR5, RBBP5 and MLL1 (or the corresponding MLL3) subunits, which accounts for the opposite roles of WDR5 in regulating the activity of the two enzymes. These findings transform our understanding of the structural basis for the regulation of MLL activity at the nucleosome level, and highlight the pivotal role of nucleosome regulation in histone-tail modification. 履歴 登録 2019年7月20日 - ヘッダ(付随情報) 公開 2019年9月11日 - マップ公開 2019年9月11日 - 更新 2024年3月27日 - 現状 2024年3月27日 処理サイト : PDBj / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト) /
Homo sapiens (ヒト) /  データ登録者
データ登録者 中国, 1件
中国, 1件  引用
引用 ジャーナル: Nature / 年: 2019
ジャーナル: Nature / 年: 2019
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_0694.map.gz
emd_0694.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-0694-v30.xml
emd-0694-v30.xml emd-0694.xml
emd-0694.xml EMDBヘッダ
EMDBヘッダ emd_0694_fsc.xml
emd_0694_fsc.xml FSCデータファイル
FSCデータファイル emd_0694.png
emd_0694.png emd_0694_msk_1.map
emd_0694_msk_1.map マスクマップ
マスクマップ emd-0694.cif.gz
emd-0694.cif.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-0694
http://ftp.pdbj.org/pub/emdb/structures/EMD-0694 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0694
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0694 emd_0694_validation.pdf.gz
emd_0694_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_0694_full_validation.pdf.gz
emd_0694_full_validation.pdf.gz emd_0694_validation.xml.gz
emd_0694_validation.xml.gz emd_0694_validation.cif.gz
emd_0694_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0694
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0694 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0694
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0694 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_0694.map.gz / 形式: CCP4 / 大きさ: 75.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_0694.map.gz / 形式: CCP4 / 大きさ: 75.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) emd_0694_msk_1.map
emd_0694_msk_1.map 試料の構成要素
試料の構成要素 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー






















 Z
Z Y
Y X
X