[English] 日本語
 Yorodumi
Yorodumi- SASDAV3: Geminin:Cdt1 2:1 heterotrimer (Geminin + DNA replication factor C... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDAV3 |
|---|---|
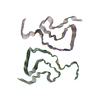
 Sample Sample | Geminin:Cdt1 2:1 heterotrimer
|
| Function / homology |  Function and homology information Function and homology informationDNA replication preinitiation complex assembly / response to sorbitol / positive regulation of DNA-templated DNA replication / regulation of nuclear cell cycle DNA replication / Switching of origins to a post-replicative state / negative regulation of DNA-templated DNA replication / attachment of mitotic spindle microtubules to kinetochore / DNA replication checkpoint signaling / positive regulation of chromatin binding / regulation of DNA-templated DNA replication initiation ...DNA replication preinitiation complex assembly / response to sorbitol / positive regulation of DNA-templated DNA replication / regulation of nuclear cell cycle DNA replication / Switching of origins to a post-replicative state / negative regulation of DNA-templated DNA replication / attachment of mitotic spindle microtubules to kinetochore / DNA replication checkpoint signaling / positive regulation of chromatin binding / regulation of DNA-templated DNA replication initiation / G1/S-Specific Transcription / negative regulation of DNA replication / negative regulation of cell cycle / regulation of DNA replication / Activation of the pre-replicative complex / DNA polymerase binding / transcription repressor complex / regulation of mitotic cell cycle / animal organ morphogenesis / positive regulation of DNA replication / Assembly of the pre-replicative complex / kinetochore / histone deacetylase binding / Orc1 removal from chromatin / transcription corepressor activity / mitotic cell cycle / DNA-binding transcription factor binding / nuclear body / cell division / negative regulation of DNA-templated transcription / chromatin binding / DNA binding / nucleoplasm / nucleus / cytosol / cytoplasm Similarity search - Function |
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2009 Journal: Proc Natl Acad Sci U S A / Year: 2009Title: Quaternary structure of the human Cdt1-Geminin complex regulates DNA replication licensing. Authors: V De Marco / P J Gillespie / A Li / N Karantzelis / E Christodoulou / R Klompmaker / S van Gerwen / A Fish / M V Petoukhov / M S Iliou / Z Lygerou / R H Medema / J J Blow / D I Svergun / S ...Authors: V De Marco / P J Gillespie / A Li / N Karantzelis / E Christodoulou / R Klompmaker / S van Gerwen / A Fish / M V Petoukhov / M S Iliou / Z Lygerou / R H Medema / J J Blow / D I Svergun / S Taraviras / A Perrakis /  Abstract: All organisms need to ensure that no DNA segments are rereplicated in a single cell cycle. Eukaryotes achieve this through a process called origin licensing, which involves tight spatiotemporal ...All organisms need to ensure that no DNA segments are rereplicated in a single cell cycle. Eukaryotes achieve this through a process called origin licensing, which involves tight spatiotemporal control of the assembly of prereplicative complexes (pre-RCs) onto chromatin. Cdt1 is a key component and crucial regulator of pre-RC assembly. In higher eukaryotes, timely inhibition of Cdt1 by Geminin is essential to prevent DNA rereplication. Here, we address the mechanism of DNA licensing inhibition by Geminin, by combining X-ray crystallography, small-angle X-ray scattering, and functional studies in Xenopus and mammalian cells. Our findings show that the Cdt1:Geminin complex can exist in two distinct forms, a "permissive" heterotrimer and an "inhibitory" heterohexamer. Specific Cdt1 residues, buried in the heterohexamer, are important for licensing. We postulate that the transition between the heterotrimer and the heterohexamer represents a molecular switch between licensing-competent and licensing-defective states. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDAV3 SASDAV3 |
|---|
-Related structure data
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
| Model #40 |  Type: atomic / Software: Crysol / Chi-square value: 3.748096  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|
- Sample
Sample
 Sample Sample | Name: Geminin:Cdt1 2:1 heterotrimer / Sample MW: 107.53 kDa / Entity id: 38 / 39 |
|---|---|
| Buffer | Name: Tris75 / Concentration: 25.00 mM / pH: 7.5 / Composition: NaCl 200.000 mM |
| Entity #38 | Type: protein / Description: Geminin / Formula weight: 23.57 / Num. of mol.: 2 / Source: Homo sapiens / References: UniProt: O75496 Sequence: MNPSMKQKQE EIKENIKNSS VPRRTLKMIQ PSASGSLVGR ENELSAGLSK RKHRNDHLTS TTSSPGVIVP ESSENKNLGG VTQESFDLMI KENPSSQYWK EVAEKRRKAL YEALKENEKL HKEIEQKDNE IARLKKENKE LAEVAEHVQY MAELIERLNG EPLDNFESLD ...Sequence: MNPSMKQKQE EIKENIKNSS VPRRTLKMIQ PSASGSLVGR ENELSAGLSK RKHRNDHLTS TTSSPGVIVP ESSENKNLGG VTQESFDLMI KENPSSQYWK EVAEKRRKAL YEALKENEKL HKEIEQKDNE IARLKKENKE LAEVAEHVQY MAELIERLNG EPLDNFESLD NQEFDSEEET VEDSLVEDSE IGTCAEGTVS SSTDAKPCI |
| Entity #39 | Name: Cdt1 / Type: protein / Description: DNA replication factor Cdt1 / Formula weight: 60.39 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: Q9H211 Sequence: MEQRRVTDFF ARRRPGPPRI APPKLACRTP SPARPALRAP ASATSGSRKR ARPPAAPGRD QARPPARRRL RLSVDEVSSP STPEAPDIPA CPSPGQKIKK STPAAGQPPH LTSAQDQDTI SELASCLQRA RELGARVRAL KASAQDAGES CTPEAEGRPE EPCGEKAPAY ...Sequence: MEQRRVTDFF ARRRPGPPRI APPKLACRTP SPARPALRAP ASATSGSRKR ARPPAAPGRD QARPPARRRL RLSVDEVSSP STPEAPDIPA CPSPGQKIKK STPAAGQPPH LTSAQDQDTI SELASCLQRA RELGARVRAL KASAQDAGES CTPEAEGRPE EPCGEKAPAY QRFHALAQPG LPGLVLPYKY QVLAEMFRSM DTIVGMLHNR SETPTFAKVQ RGVQDMMRRR FEECNVGQIK TVYPASYRFR QERSVPTFKD GTRRSDYQLT IEPLLEQEAD GAAPQLTASR LLQRRQIFSQ KLVEHVKEHH KAFLASLSPA MVVPEDQLTR WHPRFNVDEV PDIEPAALPQ PPATEKLTTA QEVLARARNL ISPRMEKALS QLALRSAAPS SPGSPRPALP ATPPATPPAA SPSALKGVSQ DLLERIRAKE AQKQLAQMTR CPEQEQRLQR LERLPELARV LRSVFVSERK PALSMEVACA RMVGSCCTIM SPGEMEKHLL LLSELLPDWL SLHRIRTDTY VKLDKAADLA HITARLAHQT RAEEGL |
-Experimental information
| Beam | Instrument name:  DORIS III X33 DORIS III X33  / City: Hamburg / 国: Germany / City: Hamburg / 国: Germany  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: MAR 345 Image Plate | |||||||||
| Scan |
| |||||||||
| Result | I0 from Guinier: 150.448 / Rg from Guinier: 2.9 nm / D max: 10 / Type of curve: single_conc /
|
 Movie
Movie Controller
Controller