[English] 日本語
 Yorodumi
Yorodumi- SASDDU9: NADPH oxidase (H2O2 producing and [F-actin] oxidizing) MICAL1 (mo... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDDU9 |
|---|---|
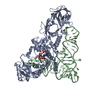
 Sample Sample | NADPH oxidase (H2O2 producing and [F-actin] oxidizing) MICAL1 (monomer) (Truncated MOCH construct)
|
| Function / homology |  Function and homology information Function and homology informationhippocampal mossy fiber expansion / F-actin monooxygenase / F-actin monooxygenase activity / NAD(P)H oxidase (H2O2-forming) / sulfur oxidation / regulation of regulated secretory pathway / NAD(P)H oxidase H2O2-forming activity / oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen / actin filament depolymerization / intermediate filament ...hippocampal mossy fiber expansion / F-actin monooxygenase / F-actin monooxygenase activity / NAD(P)H oxidase (H2O2-forming) / sulfur oxidation / regulation of regulated secretory pathway / NAD(P)H oxidase H2O2-forming activity / oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen / actin filament depolymerization / intermediate filament / actin filament bundle assembly / intercellular bridge / FAD binding / cytoskeleton organization / actin filament / monooxygenase activity / SH3 domain binding / small GTPase binding / actin filament binding / actin cytoskeleton / Factors involved in megakaryocyte development and platelet production / actin binding / midbody / endosome membrane / cilium / ciliary basal body / protein kinase binding / negative regulation of apoptotic process / signal transduction / nucleoplasm / metal ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function |
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: Protein Sci / Year: 2019 Journal: Protein Sci / Year: 2019Title: Human MICAL1: Activation by the small GTPase Rab8 and small-angle X-ray scattering studies on the oligomerization state of MICAL1 and its complex with Rab8. Authors: Alessandro Esposito / Valeria Ventura / Maxim V Petoukhov / Amrita Rai / Dmitri I Svergun / Maria A Vanoni /    Abstract: Human MICAL1 is a member of a recently discovered family of multidomain proteins that couple a FAD-containing monooxygenase-like domain to typical protein interaction domains. Growing evidence ...Human MICAL1 is a member of a recently discovered family of multidomain proteins that couple a FAD-containing monooxygenase-like domain to typical protein interaction domains. Growing evidence implicates the NADPH oxidase reaction catalyzed by the flavoprotein domain in generation of hydrogen peroxide as a second messenger in an increasing number of cell types and as a specific modulator of actin filaments stability. Several proteins of the Rab families of small GTPases are emerging as regulators of MICAL activity by binding to its C-terminal helical domain presumably shifting the equilibrium from the free - auto-inhibited - conformation to the active one. We here extend the characterization of the MICAL1-Rab8 interaction and show that indeed Rab8, in the active GTP-bound state, stabilizes the active MICAL1 conformation causing a specific four-fold increase of k of the NADPH oxidase reaction. Kinetic data and small-angle X-ray scattering (SAXS) measurements support the formation of a 1:1 complex between full-length MICAL1 and Rab8 with an apparent dissociation constant of approximately 8 μM. This finding supports the hypothesis that Rab8 is a physiological regulator of MICAL1 activity and shows how the protein region preceding the C-terminal Rab-binding domain may mask one of the Rab-binding sites detected with the isolated C-terminal fragment. SAXS-based modeling allowed us to propose the first model of the free full-length MICAL1, which is consistent with an auto-inhibited conformation in which the C-terminal region prevents catalysis by interfering with the conformational changes that are predicted to occur during the catalytic cycle. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Models
| Model #2182 |  Type: atomic / Chi-square value: 5.123  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|
- Sample
Sample
 Sample Sample | Name: NADPH oxidase (H2O2 producing and [F-actin] oxidizing) MICAL1 (monomer) (Truncated MOCH construct) Specimen concentration: 4.47 mg/ml |
|---|---|
| Buffer | Name: 50 mM sodium phosphate buffer, 5 % glycerol, 100 mM NaCl, 1 mM EDTA, 1 mM DTT pH: 7.5 / Comment: MOCH buffer |
| Entity #1195 | Name: MoCh / Type: protein / Description: [F-actin]-monooxygenase MICAL1 (MoCh) / Formula weight: 67.332 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: Q8TDZ2 Sequence: MASPTSTNPA HAHFESFLQA QLCQDVLSSF QELCGALGLE PGGGLPQYHK IKDQLNYWSA KSLWTKLDKR AGQPVYQQGR ACTSTKCLVV GAGPCGLRVA VELALLGARV VLVEKRTKFS RHNVLHLWPF TIHDLRALGA KKFYGRFCTG TLDHISIRQL QLLLLKVALL ...Sequence: MASPTSTNPA HAHFESFLQA QLCQDVLSSF QELCGALGLE PGGGLPQYHK IKDQLNYWSA KSLWTKLDKR AGQPVYQQGR ACTSTKCLVV GAGPCGLRVA VELALLGARV VLVEKRTKFS RHNVLHLWPF TIHDLRALGA KKFYGRFCTG TLDHISIRQL QLLLLKVALL LGVEIHWGVT FTGLQPPPRK GSGWRAQLQP NPPAQLANYE FDVLISAAGG KFVPEGFKVR EMRGKLAIGI TANFVNGRTV EETQVPEISG VARIYNQSFF QSLLKATGID LENIVYYKDD THYFVMTAKK QCLLRLGVLR QDWPDTNRLL GSANVVPEAL QRFTRAAADF ATHGKLGKLE FAQDAHGQPD VSAFDFTSMM RAESSARVQE KHGARLLLGL VGDCLVEPFW PLGTGVARGF LAAFDAAWMV KRWAEGAESL EVLAERESLY QLLSQTSPEN MHRNVAQYGL DPATRYPNLN LRAVTPNQVR DLYDVLAKEP VQRNNDKTDT GMPATGSAGT QEELLRWCQE QTAGYPGVHV SDLSSSWADG LALCALVYRL QPGLLEPSEL QGLGALEATA WALKVAENEL GITPVVSAQA VVAGSDPLGL IAYLSHFHSA FKSM |
-Experimental information
| Beam | Instrument name: PETRA III EMBL P12 / City: Hamburg / 国: Germany  / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3 mm / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3 mm | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 2M | |||||||||||||||||||||
| Scan | Measurement date: Jun 6, 2016 / Storage temperature: 15 °C / Cell temperature: 15 °C / Exposure time: 0.05 sec. / Number of frames: 20 / Unit: 1/nm /
| |||||||||||||||||||||
| Distance distribution function P(R) |
| |||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller

 SASDDU9
SASDDU9