[English] 日本語
 Yorodumi
Yorodumi- PDB-9pcy: HIGH-RESOLUTION SOLUTION STRUCTURE OF REDUCED FRENCH BEAN PLASTOC... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 9pcy | ||||||
|---|---|---|---|---|---|---|---|
| Title | HIGH-RESOLUTION SOLUTION STRUCTURE OF REDUCED FRENCH BEAN PLASTOCYANIN AND COMPARISON WITH THE CRYSTAL STRUCTURE OF POPLAR PLASTOCYANIN | ||||||
 Components Components | PLASTOCYANIN | ||||||
 Keywords Keywords | ELECTRON TRANSPORT | ||||||
| Function / homology |  Function and homology information Function and homology information: / chloroplast thylakoid lumen / chloroplast thylakoid membrane / copper ion binding Similarity search - Function | ||||||
| Biological species |  Phaseolus vulgaris (common bean) Phaseolus vulgaris (common bean) | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Moore, J.M. / Lepre, C.A. / Gippert, G.P. / Chazin, W.J. / Case, D.A. / Wright, P.E. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1991 Journal: J.Mol.Biol. / Year: 1991Title: High-resolution solution structure of reduced French bean plastocyanin and comparison with the crystal structure of poplar plastocyanin. Authors: Moore, J.M. / Lepre, C.A. / Gippert, G.P. / Chazin, W.J. / Case, D.A. / Wright, P.E. #1:  Journal: Biochem.Pharm. / Year: 1990 Journal: Biochem.Pharm. / Year: 1990Title: Computational Methods for Determining Protein Structures from NMR Data Authors: Gippert, G.P. / Yip, P.F. / Wright, P.E. / Case, D.A. #2:  Journal: Science / Year: 1988 Journal: Science / Year: 1988Title: Three-Dimensional Solution Structure of Plastocyanin from the Green Alga Scenedesmus Obliquus Authors: Moore, J.M. / Case, D.A. / Chazin, W.J. / Gippert, G.P. / Havel, T.F. / Powls, R. / Wright, P.E. #3:  Journal: J.Mol.Biol. / Year: 1988 Journal: J.Mol.Biol. / Year: 1988Title: Complete Assignment of the 1H Nuclear Magnetic Resonance Spectrum of French Bean Plastocyanin. Application of an Integrated Approach to Spin System Identification in Proteins Authors: Chazin, W.J. / Rance, M. / Wright, P.E. #4:  Journal: J.Mol.Biol. / Year: 1988 Journal: J.Mol.Biol. / Year: 1988Title: Complete Assignment of the 1H Nuclear Magnetic Resonance Spectrum of French Bean Plastocyanin. Sequential Resonance Assignments, Secondary Structure and Global Fold Authors: Chazin, W.J. / Wright, P.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  9pcy.cif.gz 9pcy.cif.gz | 453.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb9pcy.ent.gz pdb9pcy.ent.gz | 376.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  9pcy.json.gz 9pcy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pc/9pcy https://data.pdbj.org/pub/pdb/validation_reports/pc/9pcy ftp://data.pdbj.org/pub/pdb/validation_reports/pc/9pcy ftp://data.pdbj.org/pub/pdb/validation_reports/pc/9pcy | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 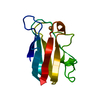
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Atom site foot note | 1: RESIDUES 16 AND 36 ARE CIS PROLINES. | |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10498.729 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Phaseolus vulgaris (common bean) / References: UniProt: P00287 Phaseolus vulgaris (common bean) / References: UniProt: P00287 |
|---|---|
| #2: Chemical | ChemComp-CU / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: other / Details: NMR |
|---|
- Processing
Processing
| Software | Name:  AMBER / Classification: refinement AMBER / Classification: refinement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| |||||||||
| NMR ensemble | Conformers submitted total number: 16 |
 Movie
Movie Controller
Controller










 PDBj
PDBj

