+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8drn | ||||||
|---|---|---|---|---|---|---|---|
| Title | LRRC8A:C conformation 1 (round) LRR focus 1 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | MEMBRANE PROTEIN / ION CHANNEL / VOLUME-REGULATION | ||||||
| Function / homology |  Function and homology information Function and homology informationMiscellaneous transport and binding events / pre-B cell differentiation / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / cell volume homeostasis / monoatomic anion transport ...Miscellaneous transport and binding events / pre-B cell differentiation / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / cell volume homeostasis / monoatomic anion transport / cellular response to osmotic stress / protein hexamerization / response to osmotic stress / monoatomic ion channel complex / positive regulation of myoblast differentiation / fat cell differentiation / intracellular glucose homeostasis / chloride transmembrane transport / electron transport chain / positive regulation of insulin secretion / spermatogenesis / periplasmic space / electron transfer activity / iron ion binding / lysosomal membrane / heme binding / endoplasmic reticulum membrane / cell surface / endoplasmic reticulum / membrane / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 4.12 Å | ||||||
 Authors Authors | Kern, D.M. / Brohawn, S.G. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structural basis for assembly and lipid-mediated gating of LRRC8A:C volume-regulated anion channels. Authors: David M Kern / Julia Bleier / Somnath Mukherjee / Jennifer M Hill / Anthony A Kossiakoff / Ehud Y Isacoff / Stephen G Brohawn /  Abstract: Leucine-rich repeat-containing protein 8 (LRRC8) family members form volume-regulated anion channels activated by hypoosmotic cell swelling. LRRC8 channels are ubiquitously expressed in vertebrate ...Leucine-rich repeat-containing protein 8 (LRRC8) family members form volume-regulated anion channels activated by hypoosmotic cell swelling. LRRC8 channels are ubiquitously expressed in vertebrate cells as heteromeric assemblies of LRRC8A (SWELL1) and LRRC8B-E subunits. Channels of different subunit composition have distinct properties that explain the functional diversity of LRRC8 currents across cell types. However, the basis for heteromeric LRRC8 channel assembly and function is unknown. Here we leverage a fiducial-tagging strategy to determine single-particle cryo-EM structures of heterohexameric LRRC8A:C channels in multiple conformations. Compared to homomers, LRRC8A:C channels show pronounced differences in architecture due to heterotypic LRR interactions that displace subunits away from the conduction axis and poise the channel for activation. Structures and functional studies further reveal that lipids embedded in the channel pore block ion conduction in the closed state. These results provide insight into determinants for heteromeric LRRC8 channel assembly, activity and gating by lipids. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8drn.cif.gz 8drn.cif.gz | 206.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8drn.ent.gz pdb8drn.ent.gz | 139 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8drn.json.gz 8drn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dr/8drn https://data.pdbj.org/pub/pdb/validation_reports/dr/8drn ftp://data.pdbj.org/pub/pdb/validation_reports/dr/8drn ftp://data.pdbj.org/pub/pdb/validation_reports/dr/8drn | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  27678MC  8dr8C  8draC  8dreC  8drkC  8droC  8drqC  8ds3C  8ds9C  8dsaC  8f74C  8f75C  8f77C  8f79C  8f7bC  8f7dC  8f7eC 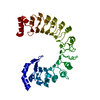 8f7jC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 105530.102 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Protein | Mass: 93624.594 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: LRRC8A:C / Type: COMPLEX / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.620 MDa / Experimental value: NO |
| Source (natural) | Organism:  |
| Source (recombinant) | Organism:  |
| Buffer solution | pH: 7.4 |
| Specimen | Conc.: 2 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: TFS KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: -2 nm / Nominal defocus min: -0.6 nm |
| Image recording | Electron dose: 50 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) |
- Processing
Processing
| Software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | |||||||||
| 3D reconstruction | Resolution: 4.12 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 110018 / Symmetry type: POINT | |||||||||
| Refinement | Cross valid method: NONE |
 Movie
Movie Controller
Controller




















 PDBj
PDBj










