+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | LRRC8A(T48D):C conformation 2 top focus | |||||||||
 Map data Map data | LRRC8A(T48D):C conformation 1 top focus | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ion channel / volume-regulation / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationMiscellaneous transport and binding events / pre-B cell differentiation / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / cell volume homeostasis / monoatomic anion transport ...Miscellaneous transport and binding events / pre-B cell differentiation / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / cell volume homeostasis / monoatomic anion transport / cellular response to osmotic stress / protein hexamerization / response to osmotic stress / monoatomic ion channel complex / positive regulation of myoblast differentiation / fat cell differentiation / intracellular glucose homeostasis / chloride transmembrane transport / electron transport chain / positive regulation of insulin secretion / spermatogenesis / periplasmic space / electron transfer activity / iron ion binding / lysosomal membrane / heme binding / endoplasmic reticulum membrane / cell surface / endoplasmic reticulum / membrane / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Kern DM / Brohawn SG | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structural basis for assembly and lipid-mediated gating of LRRC8A:C volume-regulated anion channels. Authors: David M Kern / Julia Bleier / Somnath Mukherjee / Jennifer M Hill / Anthony A Kossiakoff / Ehud Y Isacoff / Stephen G Brohawn /  Abstract: Leucine-rich repeat-containing protein 8 (LRRC8) family members form volume-regulated anion channels activated by hypoosmotic cell swelling. LRRC8 channels are ubiquitously expressed in vertebrate ...Leucine-rich repeat-containing protein 8 (LRRC8) family members form volume-regulated anion channels activated by hypoosmotic cell swelling. LRRC8 channels are ubiquitously expressed in vertebrate cells as heteromeric assemblies of LRRC8A (SWELL1) and LRRC8B-E subunits. Channels of different subunit composition have distinct properties that explain the functional diversity of LRRC8 currents across cell types. However, the basis for heteromeric LRRC8 channel assembly and function is unknown. Here we leverage a fiducial-tagging strategy to determine single-particle cryo-EM structures of heterohexameric LRRC8A:C channels in multiple conformations. Compared to homomers, LRRC8A:C channels show pronounced differences in architecture due to heterotypic LRR interactions that displace subunits away from the conduction axis and poise the channel for activation. Structures and functional studies further reveal that lipids embedded in the channel pore block ion conduction in the closed state. These results provide insight into determinants for heteromeric LRRC8 channel assembly, activity and gating by lipids. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28901.map.gz emd_28901.map.gz | 259.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28901-v30.xml emd-28901-v30.xml emd-28901.xml emd-28901.xml | 20.3 KB 20.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28901.png emd_28901.png | 45.9 KB | ||
| Others |  emd_28901_additional_1.map.gz emd_28901_additional_1.map.gz emd_28901_half_map_1.map.gz emd_28901_half_map_1.map.gz emd_28901_half_map_2.map.gz emd_28901_half_map_2.map.gz | 138.2 MB 254.4 MB 254.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28901 http://ftp.pdbj.org/pub/emdb/structures/EMD-28901 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28901 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28901 | HTTPS FTP |
-Related structure data
| Related structure data |  8f7bMC  8dr8C  8draC  8dreC  8drkC  8drnC  8droC  8drqC  8ds3C  8ds9C  8dsaC  8f74C  8f75C  8f77C  8f79C  8f7dC  8f7eC  8f7jC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28901.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28901.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LRRC8A(T48D):C conformation 1 top focus | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.048 Å | ||||||||||||||||||||||||||||||||||||
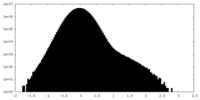
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: LRRC8A(T48D):C conformation 1 top focus unsharpened
| File | emd_28901_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LRRC8A(T48D):C conformation 1 top focus unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
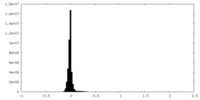
| Density Histograms |
-Half map: LRRC8A(T48D):C conformation 1 top focus
| File | emd_28901_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LRRC8A(T48D):C conformation 1 top focus | ||||||||||||
| Projections & Slices |
| ||||||||||||
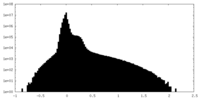
| Density Histograms |
-Half map: LRRC8A(T48D):C conformation 1 top focus
| File | emd_28901_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LRRC8A(T48D):C conformation 1 top focus | ||||||||||||
| Projections & Slices |
| ||||||||||||
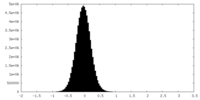
| Density Histograms |
- Sample components
Sample components
-Entire : LRRC8A:C
| Entire | Name: LRRC8A:C |
|---|---|
| Components |
|
-Supramolecule #1: LRRC8A:C
| Supramolecule | Name: LRRC8A:C / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 620 KDa |
-Macromolecule #1: Volume-regulated anion channel subunit LRRC8A,Soluble cytochrome b562
| Macromolecule | Name: Volume-regulated anion channel subunit LRRC8A,Soluble cytochrome b562 type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 105.544086 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIPVTELRYF ADTQPAYRIL KPWWDVFTDY ISIVMLMIAV FGGTLQVDQD KMICLPCKWV TKDSCNDSFR GWAASSADLE DNWETLNDN LKVIEKADNA AQVKDALTKM RAAALDAQKA TPPKLEDKSP DSPEMKDFRH GFDILVGQID DALKLANEGK V KEAQAAAE ...String: MIPVTELRYF ADTQPAYRIL KPWWDVFTDY ISIVMLMIAV FGGTLQVDQD KMICLPCKWV TKDSCNDSFR GWAASSADLE DNWETLNDN LKVIEKADNA AQVKDALTKM RAAALDAQKA TPPKLEDKSP DSPEMKDFRH GFDILVGQID DALKLANEGK V KEAQAAAE QLKTTRNAYI QKYLDTGPTG IKYDLDRHQY NYVDAVCYEN RLHWFAKYFP YLVLLHTLIF LACSNFWFKF PR TSSKLEH FVSILLKCFD SPWTTRALSE TVVEESDPKP AFSKMNGSMD KKSSTVSEDV EATVPMLQRT KSRIEQGIVD RSE TGVLDK KEGEQAKALF EKVKKFRTHV EEGDIVYRLY MRQTIIKVIK FALIICYTVY YVHNIKFDVD CTVDIESLTG YRTY RCAHP LATLFKILAS FYISLVIFYG LICMYTLWWM LRRSLKKYSF ESIREESSYS DIPDVKNDFA FMLHLIDQYD PLYSK RFAV FLSEVSENKL RQLNLNNEWT LDKLRQRLTK NAQDKLELHL FMLSGIPDTV FDLVELEVLK LELIPDVTIP PSIAQL TGL KELWLYHTAA KIEAPALAFL RENLRALHIK FTDIKEIPLW IYSLKTLEEL HLTGNLSAEN NRYIVIDGLR ELKRLKV LR LKSNLSKLPQ VVTDVGVHLQ KLSINNEGTK LIVLNSLKKM VNLTELELIR CDLERIPHSI FSLHNLQEID LKDNNLKT I EEIISFQHLH RLTCLKLWYN HIAYIPIQIG NLTNLERLYL NRNKIEKIPT QLFYCRKLRY LDLSHNNLTF LPADIGLLQ NLQNLAVTAN RIEALPPELF QCRKLRALHL GNNVLQSLPS RVGELTNLTQ IELRGNRLEC LPVELGECPL LKRSGLVVEE DLFSTLPPE VKERLWRADK EQASNSLEVL FQ |
-Macromolecule #2: Volume-regulated anion channel subunit LRRC8C
| Macromolecule | Name: Volume-regulated anion channel subunit LRRC8C / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 93.624594 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIPVTEFRQF SEQQPAFRVL KPWWDVFTDY LSVAMLMIGV FGCTLQVMQD KIICLPKRVQ PAQNHSSVPN VSQAVISTTP LPPPKPSPT NPATVEMKGL KTDLDLQQYS FINQMCYERA LHWYAKYFPY LVLIHTLVFM LCSNFWFKFP GSSSKIEHFI S ILGKCFDS ...String: MIPVTEFRQF SEQQPAFRVL KPWWDVFTDY LSVAMLMIGV FGCTLQVMQD KIICLPKRVQ PAQNHSSVPN VSQAVISTTP LPPPKPSPT NPATVEMKGL KTDLDLQQYS FINQMCYERA LHWYAKYFPY LVLIHTLVFM LCSNFWFKFP GSSSKIEHFI S ILGKCFDS PWTTRALSEV SGEDSEEKDN RKNNMNRSGT IQSGPEGNLV RSQSLKSIPE KFVVDKSAAG ALDKKEGEQA KA LFEKVKK FRLHVEEGDI LYAMYVRQTV LKVIKFLIII AYNSALVSKV QFTVDCNVDI QDMTGYKNFS CNHTMAHLFS KLS FCYLCF VSIYGLTCLY TLYWLFYRSL REYSFEYVRQ ETGIDDIPDV KNDFAFMLHM IDQYDPLYSK RFAVFLSEVS ENKL KQLNL NNEWTPDKLR QKLQTNAHNR LELPLIMLSG LPDTVFEITE LQSLKLEIIK NVMIPATIAQ LDNLQELCLH QCSVK IHSA ALSFLKENLK VLSVKFDDMR ELPPWMYGLR NLEELYLVGS LSHDISKNVT LESLRDLKSL KILSIKSNVS KIPQAV VDV SSHLQKMCVH NDGTKLVMLN NLKKMTNLTE LELVHCDLER IPHAVFSLLS LQELDLKENN LKSIEEIVSF QHLRKLT VL KLWYNSIAYI PEHIKKLTSL ERLFFSHNKV EVLPSHLFLC NKIRYLDLSY NDIRFIPPEI GVLQSLQYFS ITCNKVES L PDELYFCKKL KTLKIGKNSL SVLSPKIGNL LFLSYLDIKG NHFEVLPPEL GDCRALKRAG LVVEDALFET LPSDVREQM KADSNSENLY FQG |
-Macromolecule #3: 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine
| Macromolecule | Name: 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine / type: ligand / ID: 3 / Number of copies: 17 / Formula: PEE |
|---|---|
| Molecular weight | Theoretical: 744.034 Da |
| Chemical component information |  ChemComp-PEE: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 6 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.15 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 71566 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)