[English] 日本語
 Yorodumi
Yorodumi- PDB-7zvf: Crystal structure of human cathepsin L in complex with covalently... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7zvf | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of human cathepsin L in complex with covalently bound CLIK148 | ||||||||||||||||||
 Components Components | Cathepsin L | ||||||||||||||||||
 Keywords Keywords | HYDROLASE / cystein protease / drug target / lysosome / virus cell entry | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationenkephalin processing / cathepsin L / CD4-positive, alpha-beta T cell lineage commitment / macrophage apoptotic process / chromaffin granule / antigen processing and presentation of peptide antigen / elastin catabolic process / HS-GAG degradation / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / endolysosome lumen ...enkephalin processing / cathepsin L / CD4-positive, alpha-beta T cell lineage commitment / macrophage apoptotic process / chromaffin granule / antigen processing and presentation of peptide antigen / elastin catabolic process / HS-GAG degradation / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / endolysosome lumen / cellular response to thyroid hormone stimulus / Trafficking and processing of endosomal TLR / proteoglycan binding / Assembly of collagen fibrils and other multimeric structures / zymogen activation / antigen processing and presentation / Collagen degradation / protein autoprocessing / collagen catabolic process / fibronectin binding / serpin family protein binding / collagen binding / Degradation of the extracellular matrix / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / multivesicular body / endocytic vesicle lumen / cysteine-type peptidase activity / MHC class II antigen presentation / lysosomal lumen / : / Endosomal/Vacuolar pathway / Degradation of CDH1 / antigen processing and presentation of exogenous peptide antigen via MHC class II / extracellular matrix / histone binding / adaptive immune response / Attachment and Entry / lysosome / apical plasma membrane / fusion of virus membrane with host plasma membrane / cysteine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / symbiont entry into host cell / Golgi apparatus / proteolysis / : / extracellular exosome / extracellular region / nucleus / plasma membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.6 Å MOLECULAR REPLACEMENT / Resolution: 1.6 Å | ||||||||||||||||||
 Authors Authors | Falke, S. / Lieske, J. / Guenther, S. / Reinke, P.Y.A. / Ewert, W. / Loboda, J. / Karnicar, K. / Usenik, A. / Lindic, N. / Sekirnik, A. ...Falke, S. / Lieske, J. / Guenther, S. / Reinke, P.Y.A. / Ewert, W. / Loboda, J. / Karnicar, K. / Usenik, A. / Lindic, N. / Sekirnik, A. / Tsuge, H. / Chapman, H.N. / Hinrichs, W. / Turk, D. / Meents, A. | ||||||||||||||||||
| Funding support |  Germany, Germany,  Slovenia, 5items Slovenia, 5items
| ||||||||||||||||||
 Citation Citation |  Journal: J.Med.Chem. / Year: 2024 Journal: J.Med.Chem. / Year: 2024Title: Structural Elucidation and Antiviral Activity of Covalent Cathepsin L Inhibitors. Authors: Falke, S. / Lieske, J. / Herrmann, A. / Loboda, J. / Karnicar, K. / Gunther, S. / Reinke, P.Y.A. / Ewert, W. / Usenik, A. / Lindic, N. / Sekirnik, A. / Dretnik, K. / Tsuge, H. / Turk, V. / ...Authors: Falke, S. / Lieske, J. / Herrmann, A. / Loboda, J. / Karnicar, K. / Gunther, S. / Reinke, P.Y.A. / Ewert, W. / Usenik, A. / Lindic, N. / Sekirnik, A. / Dretnik, K. / Tsuge, H. / Turk, V. / Chapman, H.N. / Hinrichs, W. / Ebert, G. / Turk, D. / Meents, A. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7zvf.cif.gz 7zvf.cif.gz | 343.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7zvf.ent.gz pdb7zvf.ent.gz | 282.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7zvf.json.gz 7zvf.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zv/7zvf https://data.pdbj.org/pub/pdb/validation_reports/zv/7zvf ftp://data.pdbj.org/pub/pdb/validation_reports/zv/7zvf ftp://data.pdbj.org/pub/pdb/validation_reports/zv/7zvf | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 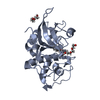 7qkdC  7zs7C  7zxaC  8a4uC  8a4vC  8a4wC  8a4xC  8a5bC  8ahvC  8b4fC  8c77C  8ofaC  8ozaC  8prxC  8qkbC  3of9S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| 2 | 
| ||||||||||||
| 3 | 
| ||||||||||||
| 4 | 
| ||||||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 4 molecules ABCD
| #1: Protein | Mass: 24161.676 Da / Num. of mol.: 4 / Mutation: T110A Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: CTSL, CTSL1 / Production host: Homo sapiens (human) / Gene: CTSL, CTSL1 / Production host:  Komagataella phaffii GS115 (fungus) / References: UniProt: P07711 Komagataella phaffii GS115 (fungus) / References: UniProt: P07711 |
|---|
-Non-polymers , 8 types, 524 molecules 














| #2: Chemical | ChemComp-KXL / ( #3: Chemical | ChemComp-PG4 / | #4: Chemical | ChemComp-1PE / #5: Chemical | ChemComp-DMS / #6: Chemical | ChemComp-PEG / #7: Chemical | #8: Chemical | ChemComp-EDO / | #9: Water | ChemComp-HOH / | |
|---|
-Details
| Has ligand of interest | Y |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.13 Å3/Da / Density % sol: 42.29 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 4 Details: Mature cathepsin L at a concentration of 7 mg/ml was equilibrated against 27% w/v PEG 8000, 1 mM TCEP and 0.1 M sodium acetate at pH 4.0. Crystals, which grew at 293 K to final size after ...Details: Mature cathepsin L at a concentration of 7 mg/ml was equilibrated against 27% w/v PEG 8000, 1 mM TCEP and 0.1 M sodium acetate at pH 4.0. Crystals, which grew at 293 K to final size after approximately 3 days, were transferred to a compound soaking solution containing 22% w/v PEG 8000, 1 mM TCEP and 0.1 M sodium acetate at pH 4.0 as well as 5% v/v DMSO and 10% v/v PEG 400. |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  PETRA III, DESY PETRA III, DESY  / Beamline: P11 / Wavelength: 1.033 Å / Beamline: P11 / Wavelength: 1.033 Å |
| Detector | Type: DECTRIS EIGER2 X 16M / Detector: PIXEL / Date: Feb 20, 2022 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.033 Å / Relative weight: 1 |
| Reflection | Resolution: 1.59→44.4 Å / Num. obs: 100813 / % possible obs: 94.2 % / Redundancy: 11 % / Biso Wilson estimate: 15.15 Å2 / CC1/2: 0.998 / Rrim(I) all: 0.17 / Net I/σ(I): 12.63 |
| Reflection shell | Resolution: 1.59→1.63 Å / Redundancy: 10.1 % / Mean I/σ(I) obs: 2.75 / Num. unique obs: 7434 / CC1/2: 0.873 / Rrim(I) all: 0.886 / % possible all: 93.4 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3OF9 Resolution: 1.6→44.33 Å / SU ML: 0.1642 / Cross valid method: FREE R-VALUE / σ(F): 1.98 / Phase error: 21.2679 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.48 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→44.33 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller


 PDBj
PDBj




















