+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7lp1 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of Nedd4L WW3 domain | ||||||
 Components Components | E3 ubiquitin-protein ligase NEDD4-like | ||||||
 Keywords Keywords | LIGASE / PPxY binding / E3 Ubiquitin ligase / Nedd4L | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of caveolin-mediated endocytosis / RING-type E3 ubiquitin transferase (cysteine targeting) / negative regulation of sodium ion transmembrane transport / negative regulation of sodium ion import across plasma membrane / negative regulation of potassium ion export across plasma membrane / negative regulation of potassium ion transmembrane transport / negative regulation of protein localization to cell surface / regulation of membrane repolarization / positive regulation of dendrite extension / regulation of membrane depolarization ...positive regulation of caveolin-mediated endocytosis / RING-type E3 ubiquitin transferase (cysteine targeting) / negative regulation of sodium ion transmembrane transport / negative regulation of sodium ion import across plasma membrane / negative regulation of potassium ion export across plasma membrane / negative regulation of potassium ion transmembrane transport / negative regulation of protein localization to cell surface / regulation of membrane repolarization / positive regulation of dendrite extension / regulation of membrane depolarization / receptor catabolic process / regulation of sodium ion transmembrane transport / potassium channel inhibitor activity / ventricular cardiac muscle cell action potential / HECT-type E3 ubiquitin transferase / sodium channel inhibitor activity / regulation of dendrite morphogenesis / neuromuscular junction development / regulation of synapse organization / sodium channel regulator activity / protein monoubiquitination / protein K48-linked ubiquitination / multivesicular body / Downregulation of TGF-beta receptor signaling / regulation of membrane potential / Downregulation of SMAD2/3:SMAD4 transcriptional activity / Budding and maturation of HIV virion / regulation of protein stability / receptor internalization / Stimuli-sensing channels / neuron projection development / ubiquitin-protein transferase activity / positive regulation of protein catabolic process / ubiquitin protein ligase activity / Antigen processing: Ubiquitination & Proteasome degradation / ubiquitin-dependent protein catabolic process / monoatomic ion transmembrane transport / proteasome-mediated ubiquitin-dependent protein catabolic process / transmembrane transporter binding / protein ubiquitination / apical plasma membrane / Golgi apparatus / extracellular exosome / nucleoplasm / cytosol / cytoplasm Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.35 Å MOLECULAR REPLACEMENT / Resolution: 1.35 Å | ||||||
 Authors Authors | Alian, A. / Alam, S.L. / Thompson, T. / Rheinemann, L. / Sundquist, W.I. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2021 Journal: J.Biol.Chem. / Year: 2021Title: Interactions between AMOT PPxY motifs and NEDD4L WW domains function in HIV-1 release. Authors: Rheinemann, L. / Thompson, T. / Mercenne, G. / Paine, E.L. / Peterson, F.C. / Volkman, B.F. / Alam, S.L. / Alian, A. / Sundquist, W.I. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7lp1.cif.gz 7lp1.cif.gz | 46.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7lp1.ent.gz pdb7lp1.ent.gz | 26.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7lp1.json.gz 7lp1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7lp1_validation.pdf.gz 7lp1_validation.pdf.gz | 428.1 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7lp1_full_validation.pdf.gz 7lp1_full_validation.pdf.gz | 428.1 KB | Display | |
| Data in XML |  7lp1_validation.xml.gz 7lp1_validation.xml.gz | 4.2 KB | Display | |
| Data in CIF |  7lp1_validation.cif.gz 7lp1_validation.cif.gz | 5 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lp/7lp1 https://data.pdbj.org/pub/pdb/validation_reports/lp/7lp1 ftp://data.pdbj.org/pub/pdb/validation_reports/lp/7lp1 ftp://data.pdbj.org/pub/pdb/validation_reports/lp/7lp1 | HTTPS FTP |
-Related structure data
| Related structure data |  7lp2C 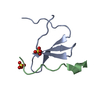 7lp3C  7lp4C  7lp5C  2mptS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 4638.264 Da / Num. of mol.: 1 / Fragment: WW 3 domain, residues 494-532 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: NEDD4L, KIAA0439, NEDL3 Homo sapiens (human) / Gene: NEDD4L, KIAA0439, NEDL3Production host:  References: UniProt: Q96PU5, HECT-type E3 ubiquitin transferase | ||||||
|---|---|---|---|---|---|---|---|
| #2: Chemical | | #3: Chemical | #4: Water | ChemComp-HOH / | Has ligand of interest | N | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.78 Å3/Da / Density % sol: 55.73 % |
|---|---|
| Crystal grow | Temperature: 293.15 K / Method: vapor diffusion, sitting drop / pH: 4.6 Details: 3.6M ammonium nitrate, 5% glycerol, and 0.1M sodium acetate at pH 4.6 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL7-1 / Wavelength: 1.1 Å / Beamline: BL7-1 / Wavelength: 1.1 Å |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Jun 17, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.1 Å / Relative weight: 1 |
| Reflection | Resolution: 1→50 Å / Num. obs: 11774 / % possible obs: 99.83 % / Redundancy: 2 % / Biso Wilson estimate: 13 Å2 / CC1/2: 1 / Net I/σ(I): 33.25 |
| Reflection shell | Resolution: 1.35→1.398 Å / Redundancy: 2 % / Rmerge(I) obs: 0.1685 / Mean I/σ(I) obs: 4.26 / Num. unique obs: 1140 / CC1/2: 0.958 / Rrim(I) all: 0.2383 / % possible all: 98.16 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2mpt Resolution: 1.35→29.47 Å / SU ML: 0.095 / Cross valid method: FREE R-VALUE / σ(F): 1.34 / Phase error: 23.5887 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 20.39 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.35→29.47 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Refine-ID: X-RAY DIFFRACTION / Auth asym-ID: A / Label asym-ID: A
|
 Movie
Movie Controller
Controller










 PDBj
PDBj











