+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7dac | ||||||
|---|---|---|---|---|---|---|---|
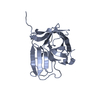
| Title | Human RIPK3 amyloid fibril revealed by solid-state NMR | ||||||
 Components Components | Receptor-interacting serine/threonine-protein kinase 3 | ||||||
 Keywords Keywords | PROTEIN FIBRIL / programmed necrosis / amyloid | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of activation-induced cell death of T cells / regulation of CD8-positive, alpha-beta cytotoxic T cell extravasation / execution phase of necroptosis / regulation of T cell mediated cytotoxicity / ripoptosome assembly involved in necroptotic process / regulation of adaptive immune response / regulation of activated T cell proliferation / activation of protein kinase activity / Microbial modulation of RIPK1-mediated regulated necrosis / ripoptosome ...regulation of activation-induced cell death of T cells / regulation of CD8-positive, alpha-beta cytotoxic T cell extravasation / execution phase of necroptosis / regulation of T cell mediated cytotoxicity / ripoptosome assembly involved in necroptotic process / regulation of adaptive immune response / regulation of activated T cell proliferation / activation of protein kinase activity / Microbial modulation of RIPK1-mediated regulated necrosis / ripoptosome / TRIF-mediated programmed cell death / regulation of type II interferon production / TLR3-mediated TICAM1-dependent programmed cell death / programmed necrotic cell death / SARS-CoV-1-mediated effects on programmed cell death / necroptotic signaling pathway / RIP-mediated NFkB activation via ZBP1 / positive regulation of necroptotic process / RIPK1-mediated regulated necrosis / non-canonical NF-kappaB signal transduction / TRP channels / necroptotic process / T cell homeostasis / lymph node development / spleen development / positive regulation of intrinsic apoptotic signaling pathway / reactive oxygen species metabolic process / thymus development / protein modification process / TICAM1, RIP1-mediated IKK complex recruitment / : / protein serine/threonine kinase binding / IKK complex recruitment mediated by RIP1 / apoptotic signaling pathway / Regulation of necroptotic cell death / positive regulation of reactive oxygen species metabolic process / cellular response to hydrogen peroxide / SARS-CoV-1 activates/modulates innate immune responses / T cell differentiation in thymus / regulation of apoptotic process / amyloid fibril formation / defense response to virus / protein kinase activity / transcription coactivator activity / non-specific serine/threonine protein kinase / protein serine kinase activity / protein serine/threonine kinase activity / protein-containing complex binding / signal transduction / protein-containing complex / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLID-STATE NMR / simulated annealing | ||||||
 Authors Authors | Wu, X.L. / Zhang, J. / Dong, X.Q. / Liu, J. / Li, B. / Hu, H. / Wang, J. / Wang, H.Y. / Lu, J.X. | ||||||
| Funding support |  China, 1items China, 1items
| ||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2021 Journal: Proc Natl Acad Sci U S A / Year: 2021Title: The structure of a minimum amyloid fibril core formed by necroptosis-mediating RHIM of human RIPK3. Authors: Xialian Wu / Yeyang Ma / Kun Zhao / Jing Zhang / Yunpeng Sun / Yichen Li / Xingqi Dong / Hong Hu / Jing Liu / Jian Wang / Xia Zhang / Bing Li / Huayi Wang / Dan Li / Bo Sun / Junxia Lu / Cong Liu /  Abstract: Receptor-interacting protein kinases 3 (RIPK3), a central node in necroptosis, polymerizes in response to the upstream signals and then activates its downstream mediator to induce cell death. The ...Receptor-interacting protein kinases 3 (RIPK3), a central node in necroptosis, polymerizes in response to the upstream signals and then activates its downstream mediator to induce cell death. The active polymeric form of RIPK3 has been indicated as the form of amyloid fibrils assembled via its RIP homotypic interaction motif (RHIM). In this study, we combine cryogenic electron microscopy and solid-state NMR to determine the amyloid fibril structure of RIPK3 RHIM-containing C-terminal domain (CTD). The structure reveals a single protofilament composed of the RHIM domain. RHIM forms three β-strands (referred to as strands 1 through 3) folding into an S shape, a distinct fold from that in complex with RIPK1. The consensus tetrapeptide VQVG of RHIM forms strand 2, which zips up strands 1 and 3 via heterozipper-like interfaces. Notably, the RIPK3-CTD fibril, as a physiological fibril, exhibits distinctive assembly compared with pathological fibrils. It has an exceptionally small fibril core and twists in both handedness with the smallest pitch known so far. These traits may contribute to a favorable spatial arrangement of RIPK3 kinase domain for efficient phosphorylation. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7dac.cif.gz 7dac.cif.gz | 551.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7dac.ent.gz pdb7dac.ent.gz | 436.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7dac.json.gz 7dac.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/da/7dac https://data.pdbj.org/pub/pdb/validation_reports/da/7dac ftp://data.pdbj.org/pub/pdb/validation_reports/da/7dac ftp://data.pdbj.org/pub/pdb/validation_reports/da/7dac | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7da4C C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11863.072 Da / Num. of mol.: 5 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: RIPK3, RIP3 / Production host: Homo sapiens (human) / Gene: RIPK3, RIP3 / Production host:  References: UniProt: Q9Y572, non-specific serine/threonine protein kinase |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLID-STATE NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj






 microscopy
microscopy