[English] 日本語
 Yorodumi
Yorodumi- PDB-6xnf: GCN4-p1 Peptide Trimer with Tetrafluoroiodophenylalanine residue ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6xnf | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GCN4-p1 Peptide Trimer with Tetrafluoroiodophenylalanine residue at position 16 (TFI-F16) | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | TRANSCRIPTION / Coil-coil / tetrafluoroiodo-modified phenylalanine | |||||||||
| Function / homology |  Function and homology information Function and homology informationFCERI mediated MAPK activation / protein localization to nuclear periphery / Activation of the AP-1 family of transcription factors / negative regulation of ribosomal protein gene transcription by RNA polymerase II / positive regulation of cellular response to amino acid starvation / response to amino acid starvation / mediator complex binding / Oxidative Stress Induced Senescence / amino acid biosynthetic process / TFIID-class transcription factor complex binding ...FCERI mediated MAPK activation / protein localization to nuclear periphery / Activation of the AP-1 family of transcription factors / negative regulation of ribosomal protein gene transcription by RNA polymerase II / positive regulation of cellular response to amino acid starvation / response to amino acid starvation / mediator complex binding / Oxidative Stress Induced Senescence / amino acid biosynthetic process / TFIID-class transcription factor complex binding / positive regulation of RNA polymerase II transcription preinitiation complex assembly / positive regulation of transcription initiation by RNA polymerase II / cellular response to nutrient levels / cellular response to amino acid starvation / RNA polymerase II transcription regulator complex / DNA-binding transcription activator activity, RNA polymerase II-specific / transcription regulator complex / sequence-specific DNA binding / RNA polymerase II-specific DNA-binding transcription factor binding / DNA-binding transcription factor activity, RNA polymerase II-specific / intracellular signal transduction / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / chromatin binding / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / identical protein binding / nucleus Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | |||||||||
 Authors Authors | Rowe Hartje, R.K. / Czarny, R.S. / Ho, A. | |||||||||
| Funding support |  United States, 2items United States, 2items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Engineering Specific Protein-Protein Interactions Through Halogen and Hydrogen Bonds Authors: Rowe Hartje, R.K. / Ferrero, M. / Cavallo, G. / Metrangolo, P. / Ho, A. / Czarny, R. / Ho, P.S. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6xnf.cif.gz 6xnf.cif.gz | 64.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6xnf.ent.gz pdb6xnf.ent.gz | 40.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6xnf.json.gz 6xnf.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xn/6xnf https://data.pdbj.org/pub/pdb/validation_reports/xn/6xnf ftp://data.pdbj.org/pub/pdb/validation_reports/xn/6xnf ftp://data.pdbj.org/pub/pdb/validation_reports/xn/6xnf | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6xneC  6xnlC  6xnmC  1swiS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 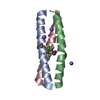
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 3893.234 Da / Num. of mol.: 1 / Mutation: TFI-F16 / Source method: obtained synthetically / Source: (synth.)  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 3619.280 Da / Num. of mol.: 2 / Mutation: TFI-F16 / Source method: obtained synthetically / Source: (synth.)  #3: Chemical | ChemComp-NA / #4: Water | ChemComp-HOH / | Has ligand of interest | Y | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.18 Å3/Da / Density % sol: 43.54 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7 Details: Crystallization drops were prepared by mixing 2 uL of stock peptide solution with 2 uL of mother liquor and allowed to equilibrate at 298 K over a well containing 500 uL of mother liquor. ...Details: Crystallization drops were prepared by mixing 2 uL of stock peptide solution with 2 uL of mother liquor and allowed to equilibrate at 298 K over a well containing 500 uL of mother liquor. The stock peptide solution (total concentration 1.5 mM) was prepared by mixing 2:1 ratios of the A16 peptide with TFI-F16 in 10 mM potassium phosphate, 100 mM potassium chloride pH 7.0. |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU MICROMAX-003 / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU MICROMAX-003 / Wavelength: 1.5418 Å |
| Detector | Type: DECTRIS PILATUS 200K / Detector: PIXEL / Date: Aug 8, 2017 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2→46.21 Å / Num. obs: 43081 / % possible obs: 99.47 % / Redundancy: 6.5 % / Biso Wilson estimate: 28.81 Å2 / CC1/2: 0.996 / CC star: 0.999 / Rmerge(I) obs: 0.09809 / Rpim(I) all: 0.04159 / Rrim(I) all: 0.1068 / Net I/σ(I): 11.38 |
| Reflection shell | Resolution: 2→2.072 Å / Redundancy: 6 % / Rmerge(I) obs: 0.7578 / Mean I/σ(I) obs: 2.69 / Num. unique obs: 3998 / CC1/2: 0.766 / CC star: 0.931 / Rpim(I) all: 0.8326 / Rrim(I) all: 0.3375 / % possible all: 99.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1SWI Resolution: 2→46.21 Å / SU ML: 0.2044 / Cross valid method: FREE R-VALUE / σ(F): 1.34 / Phase error: 30.4397 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 39.45 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→46.21 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj




