+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 6xf5 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
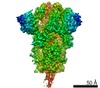
| タイトル | Cryo-EM structure of a biotinylated SARS-CoV-2 spike probe in the prefusion state (RBDs down) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 要素 要素 | Spike glycoprotein | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 キーワード キーワード | VIRAL PROTEIN / Fusion protein / Molecular probe / Spike glycoprotein / COVID-19 / RBD | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / viral translation / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / viral translation / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / membrane fusion / Attachment and Entry / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / receptor ligand activity / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane 類似検索 - 分子機能 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 生物種 |  | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 手法 | 電子顕微鏡法 / 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.45 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 データ登録者 データ登録者 | Cerutti, G. / Gorman, J. / Kwong, P.D. / Shapiro, L. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 引用 引用 |  ジャーナル: SSRN / 年: 2020 ジャーナル: SSRN / 年: 2020タイトル: Structure-Based Design with Tag-Based Purification and In-Process Biotinylation Enable Streamlined Development of SARS-CoV-2 Spike Molecular Probes. 著者: Tongqing Zhou / I-Ting Teng / Adam S Olia / Gabriele Cerutti / Jason Gorman / Alexandra Nazzari / Wei Shi / Yaroslav Tsybovsky / Lingshu Wang / Shuishu Wang / Baoshan Zhang / Yi Zhang / ...著者: Tongqing Zhou / I-Ting Teng / Adam S Olia / Gabriele Cerutti / Jason Gorman / Alexandra Nazzari / Wei Shi / Yaroslav Tsybovsky / Lingshu Wang / Shuishu Wang / Baoshan Zhang / Yi Zhang / Phinikoula S Katsamba / Yuliya Petrova / Bailey B Banach / Ahmed S Fahad / Lihong Liu / Sheila N Lopez Acevedo / Bharat Madan / Matheus Olivera de Souza / Xiaoli Pan / Pengfei Wang / Jacy R Wolfe / Michael Yin / David D Ho / Emily Phung / Anthony DiPiazza / Lauren Chang / Olubukula Abiona / Kizzmekia S Corbett / Brandon J DeKosky / Barney S Graham / John R Mascola / John Misasi / Tracy Ruckwardt / Nancy J Sullivan / Lawrence Shapiro / Peter D Kwong /  要旨: Biotin-labeled molecular probes, comprising specific regions of the SARS-CoV-2 spike, would be helpful in the isolation and characterization of antibodies targeting this recently emerged pathogen. To ...Biotin-labeled molecular probes, comprising specific regions of the SARS-CoV-2 spike, would be helpful in the isolation and characterization of antibodies targeting this recently emerged pathogen. To develop such probes, we designed constructs incorporating an N-terminal purification tag, a site-specific protease-cleavage site, the probe region of interest, and a C-terminal sequence targeted by biotin ligase. Probe regions included full-length spike ectodomain as well as various subregions, and we also designed mutants to eliminate recognition of the ACE2 receptor. Yields of biotin-labeled probes from transient transfection ranged from ~0.5 mg/L for the complete ectodomain to >5 mg/L for several subregions. Probes were characterized for antigenicity and ACE2 recognition, and the structure of the spike ectodomain probe was determined by cryo-electron microscopy. We also characterized antibody-binding specificities and cell-sorting capabilities of the biotinylated probes. Altogether, structure-based design coupled to efficient purification and biotinylation processes can thus enable streamlined development of SARS-CoV-2 spike-ectodomain probes. Funding: Support for this work was provided by the Intramural Research Program of the Vaccine Research Center, National Institute of Allergy and Infectious Diseases (NIAID). Support for this work was also provided by COVID-19 Fast Grants, the Jack Ma Foundation, the Self Graduate Fellowship Program, and NIH grants DP5OD023118, R21AI143407, and R21AI144408. Some of this work was performed at the Columbia University Cryo-EM Center at the Zuckerman Institute, and some at the Simons Electron Microscopy Center (SEMC) and National Center for Cryo-EM Access and Training (NCCAT) located at the New York Structural Biology Center, supported by grants from the Simons Foundation (SF349247), NYSTAR, and the NIH National Institute of General Medical Sciences (GM103310). Conflict of Interest: The authors declare that they have no conflict of interest. Ethical Approval: Peripheral blood mononuclear cells (PBMCs) for B cell sorting were obtained from a convalescent SARS-CoV-2 patient (collected 75 days post symptom onset under an IRB approved clinical trial protocol, VRC 200 - ClinicalTrials.gov Identifier: NCT00067054) and a healthy control donor from the NIH blood bank pre-SARS-CoV-2 pandemic. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  6xf5.cif.gz 6xf5.cif.gz | 563 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb6xf5.ent.gz pdb6xf5.ent.gz | 450.4 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  6xf5.json.gz 6xf5.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  6xf5_validation.pdf.gz 6xf5_validation.pdf.gz | 3.5 MB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  6xf5_full_validation.pdf.gz 6xf5_full_validation.pdf.gz | 3.6 MB | 表示 | |
| XML形式データ |  6xf5_validation.xml.gz 6xf5_validation.xml.gz | 88.2 KB | 表示 | |
| CIF形式データ |  6xf5_validation.cif.gz 6xf5_validation.cif.gz | 133.7 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/xf/6xf5 https://data.pdbj.org/pub/pdb/validation_reports/xf/6xf5 ftp://data.pdbj.org/pub/pdb/validation_reports/xf/6xf5 ftp://data.pdbj.org/pub/pdb/validation_reports/xf/6xf5 | HTTPS FTP |
-関連構造データ
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 
|
|---|---|
| 1 |
|
- 要素
要素
| #1: タンパク質 | 分子量: 138723.297 Da / 分子数: 3 / 由来タイプ: 組換発現 由来: (組換発現)  遺伝子: S, 2 / 発現宿主:  Homo sapiens (ヒト) / 参照: UniProt: P0DTC2 Homo sapiens (ヒト) / 参照: UniProt: P0DTC2#2: 多糖 | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose #3: 糖 | ChemComp-NAG / 研究の焦点であるリガンドがあるか | Y | Has protein modification | Y | |
|---|
-実験情報
-実験
| 実験 | 手法: 電子顕微鏡法 |
|---|---|
| EM実験 | 試料の集合状態: PARTICLE / 3次元再構成法: 単粒子再構成法 |
- 試料調製
試料調製
| 構成要素 | 名称: SARS-CoV-2 spike ectodomain biotinylated probe in the prefusion state タイプ: ORGANELLE OR CELLULAR COMPONENT / Entity ID: #1 / 由来: RECOMBINANT |
|---|---|
| 由来(天然) | 生物種:  |
| 由来(組換発現) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 緩衝液 | pH: 7.4 |
| 試料 | 包埋: NO / シャドウイング: NO / 染色: NO / 凍結: YES |
| 急速凍結 | 凍結剤: ETHANE |
- 電子顕微鏡撮影
電子顕微鏡撮影
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
|---|---|
| 顕微鏡 | モデル: FEI TITAN KRIOS |
| 電子銃 | 電子線源:  FIELD EMISSION GUN / 加速電圧: 300 kV / 照射モード: OTHER FIELD EMISSION GUN / 加速電圧: 300 kV / 照射モード: OTHER |
| 電子レンズ | モード: BRIGHT FIELD |
| 撮影 | 電子線照射量: 42 e/Å2 フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) |
- 解析
解析
| ソフトウェア | 名称: PHENIX / バージョン: 1.18.2_3874: / 分類: 精密化 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EMソフトウェア |
| ||||||||||||||||||||||||
| CTF補正 | タイプ: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| 対称性 | 点対称性: C3 (3回回転対称) | ||||||||||||||||||||||||
| 3次元再構成 | 解像度: 3.45 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 粒子像の数: 43176 / 対称性のタイプ: POINT | ||||||||||||||||||||||||
| 原子モデル構築 | プロトコル: FLEXIBLE FIT | ||||||||||||||||||||||||
| 原子モデル構築 | PDB-ID: 6VXX Accession code: 6VXX / Source name: PDB / タイプ: experimental model | ||||||||||||||||||||||||
| 拘束条件 |
|
 ムービー
ムービー コントローラー
コントローラー















 PDBj
PDBj






