+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6i0o | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of human IMP dehydrogenase, isoform 2, bound to GTP | ||||||
 Components Components | Inosine-5'-monophosphate dehydrogenase 2 | ||||||
 Keywords Keywords | BIOSYNTHETIC PROTEIN / oxidoreductase / de novo guanine nucleotide biosynthesis | ||||||
| Function / homology |  Function and homology information Function and homology informationlymphocyte proliferation / 'de novo' XMP biosynthetic process / Purine ribonucleoside monophosphate biosynthesis / IMP dehydrogenase / IMP dehydrogenase activity / GMP biosynthetic process / Azathioprine ADME / peroxisomal membrane / GTP biosynthetic process / cellular response to interleukin-4 ...lymphocyte proliferation / 'de novo' XMP biosynthetic process / Purine ribonucleoside monophosphate biosynthesis / IMP dehydrogenase / IMP dehydrogenase activity / GMP biosynthetic process / Azathioprine ADME / peroxisomal membrane / GTP biosynthetic process / cellular response to interleukin-4 / circadian rhythm / secretory granule lumen / Potential therapeutics for SARS / ficolin-1-rich granule lumen / nucleotide binding / Neutrophil degranulation / DNA binding / RNA binding / extracellular exosome / extracellular region / membrane / metal ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.623 Å MOLECULAR REPLACEMENT / Resolution: 2.623 Å | ||||||
 Authors Authors | Buey, R.M. / Fernandez-Justel, D. / Revuelta, J.L. | ||||||
| Funding support |  Spain, 1items Spain, 1items
| ||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2019 Journal: J Mol Biol / Year: 2019Title: A Nucleotide-Dependent Conformational Switch Controls the Polymerization of Human IMP Dehydrogenases to Modulate their Catalytic Activity. Authors: David Fernández-Justel / Rafael Núñez / Jaime Martín-Benito / David Jimeno / Adrián González-López / Eva María Soriano / José Luis Revuelta / Rubén M Buey /  Abstract: Inosine 5'-monophosphate dehydrogenase (IMPDH) catalyzes the rate-limiting step in the de novo GTP biosynthetic pathway and plays essential roles in cell proliferation. As a clinical target, IMPDH ...Inosine 5'-monophosphate dehydrogenase (IMPDH) catalyzes the rate-limiting step in the de novo GTP biosynthetic pathway and plays essential roles in cell proliferation. As a clinical target, IMPDH has been studied for decades, but it has only been within the last years that we are starting to understand the complexity of the mechanisms of its physiological regulation. Here, we report structural and functional insights into how adenine and guanine nucleotides control a conformational switch that modulates the assembly of the two human IMPDH enzymes into cytoophidia and allosterically regulates their catalytic activity. In vitro reconstituted micron-length cytoophidia-like structures show catalytic activity comparable to unassembled IMPDH but, in turn, are more resistant to GTP/GDP allosteric inhibition. Therefore, IMPDH cytoophidia formation facilitates the accumulation of high levels of guanine nucleotides when the cell requires it. Finally, we demonstrate that most of the IMPDH retinopathy-associated mutations abrogate GTP/GDP-induced allosteric inhibition and alter cytoophidia dynamics. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6i0o.cif.gz 6i0o.cif.gz | 477.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6i0o.ent.gz pdb6i0o.ent.gz | 401.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6i0o.json.gz 6i0o.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/i0/6i0o https://data.pdbj.org/pub/pdb/validation_reports/i0/6i0o ftp://data.pdbj.org/pub/pdb/validation_reports/i0/6i0o ftp://data.pdbj.org/pub/pdb/validation_reports/i0/6i0o | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 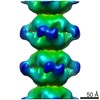 4402C 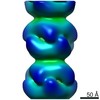 4403C  6i0mC  4z87S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 56157.137 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: IMPDH2, IMPD2 / Production host: Homo sapiens (human) / Gene: IMPDH2, IMPD2 / Production host:  #2: Chemical | ChemComp-GTP / #3: Chemical | ChemComp-SO4 / #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.26 Å3/Da / Density % sol: 62.27 % |
|---|---|
| Crystal grow | Temperature: 295 K / Method: vapor diffusion, sitting drop Details: 25% (w/v) PEG-1500 0.1 M Buffer MIB (malonic acid, imadazole, boric acid), pH 9.0 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I03 / Wavelength: 0.99987 Å / Beamline: I03 / Wavelength: 0.99987 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Sep 21, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.99987 Å / Relative weight: 1 |
| Reflection | Resolution: 2.623→81.953 Å / Num. obs: 44889 / % possible obs: 92.6 % / Redundancy: 6.5 % / CC1/2: 0.997 / Rmerge(I) obs: 0.142 / Rpim(I) all: 0.06 / Rrim(I) all: 0.155 / Net I/σ(I): 10.6 |
| Reflection shell | Resolution: 2.623→2.827 Å / Redundancy: 7 % / Rmerge(I) obs: 1.564 / Mean I/σ(I) obs: 1.5 / Num. unique obs: 353 / CC1/2: 0.408 / Rpim(I) all: 0.635 / Rrim(I) all: 1.689 / % possible all: 47 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4Z87 Resolution: 2.623→81.953 Å / SU ML: 0.32 / Cross valid method: FREE R-VALUE / σ(F): 1.35 / Phase error: 28.37 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.623→81.953 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller










 PDBj
PDBj





