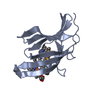
Entry Database : PDB / ID : 6g8tTitle Crystal Structures of the Single PDZ Domains from GRASP65 and their Interaction with the Golgin GM130 Golgi reassembly-stacking protein 1 Keywords / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / / Resolution : 2.67 Å Authors Jurk, C.M. / Roske, Y. / Heinemann, U. Journal : Croatica Chemica Acta / Year : 2018Title : Crystal Structures of the Single PDZ Domains from GRASP65 and their Interaction with the Golgin GM130Authors : Jurk, C.M. / Roske, Y. / Heinemann, U. History Deposition Apr 10, 2018 Deposition site / Processing site Revision 1.0 Apr 24, 2019 Provider / Type Revision 1.1 May 6, 2020 Group / Category Item _citation.country / _citation.journal_abbrev ... _citation.country / _citation.journal_abbrev / _citation.journal_id_CSD / _citation.journal_id_ISSN / _citation.pdbx_database_id_DOI / _citation.year Revision 1.2 Jan 17, 2024 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.67 Å
MOLECULAR REPLACEMENT / Resolution: 2.67 Å  Authors
Authors Citation
Citation Journal: Croatica Chemica Acta / Year: 2018
Journal: Croatica Chemica Acta / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6g8t.cif.gz
6g8t.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6g8t.ent.gz
pdb6g8t.ent.gz PDB format
PDB format 6g8t.json.gz
6g8t.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/g8/6g8t
https://data.pdbj.org/pub/pdb/validation_reports/g8/6g8t ftp://data.pdbj.org/pub/pdb/validation_reports/g8/6g8t
ftp://data.pdbj.org/pub/pdb/validation_reports/g8/6g8t


 Links
Links Assembly
Assembly
 Components
Components Homo sapiens (human) / Gene: GORASP1, GOLPH5, GRASP65 / Production host:
Homo sapiens (human) / Gene: GORASP1, GOLPH5, GRASP65 / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  BESSY
BESSY  / Beamline: 14.1 / Wavelength: 0.91841 Å
/ Beamline: 14.1 / Wavelength: 0.91841 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller











 PDBj
PDBj






