+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6d8u | ||||||
|---|---|---|---|---|---|---|---|
| Title | NMR solution structure of tamapin, mutant K20E | ||||||
 Components Components | Potassium channel toxin alpha-KTx 5.4 | ||||||
 Keywords Keywords | TOXIN / Tamapin mutant / K20E / CSalpha/beta / SK channels | ||||||
| Function / homology | Scorpion short toxins signature. / Scorpion short chain toxin, potassium channel inhibitor / Scorpion short toxin, BmKK2 / Knottin, scorpion toxin-like superfamily / ion channel inhibitor activity / potassium channel regulator activity / toxin activity / extracellular region / Potassium channel toxin alpha-KTx 5.4 Function and homology information Function and homology information | ||||||
| Biological species |  Mesobuthus tamulus (Indian red scorpion) Mesobuthus tamulus (Indian red scorpion) | ||||||
| Method | SOLUTION NMR / molecular dynamics | ||||||
 Authors Authors | del Rio Portilla, F. / Melchor Meneses, C.M. / Titaux Delgado, G.A. / Mayorga Flores, M. | ||||||
 Citation Citation |  Journal: Acs Med.Chem.Lett. / Year: 2020 Journal: Acs Med.Chem.Lett. / Year: 2020Title: Novel Blocker of Onco SK3 Channels Derived from Scorpion Toxin Tamapin and Active against Migration of Cancer Cells Authors: Flores, M. / Chantome, A. / Melchor-Meneses, C.M. / Domingo, I. / Titaux Delgado, G.A. / Galindo-Murillo, R. / Vandier, C. / del Rio Portilla, F. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6d8u.cif.gz 6d8u.cif.gz | 183.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6d8u.ent.gz pdb6d8u.ent.gz | 153.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6d8u.json.gz 6d8u.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/d8/6d8u https://data.pdbj.org/pub/pdb/validation_reports/d8/6d8u ftp://data.pdbj.org/pub/pdb/validation_reports/d8/6d8u ftp://data.pdbj.org/pub/pdb/validation_reports/d8/6d8u | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6d3tC  6d8hC  6d8qC  6d8rC  6d8sC  6d8tC  6d8yC  6d93C  6d9oC  6d9pC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 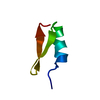
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 3470.119 Da / Num. of mol.: 1 / Mutation: K20E Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Mesobuthus tamulus (Indian red scorpion) Mesobuthus tamulus (Indian red scorpion)Production host:  |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Type: solution Contents: 2.0 mM Potassium channel toxin alpha-KTx 5.4, 95% H2O/5% D2O Label: 1H_sample / Solvent system: 95% H2O/5% D2O |
|---|---|
| Sample | Conc.: 2.0 mM / Component: Potassium channel toxin alpha-KTx 5.4 / Isotopic labeling: natural abundance |
| Sample conditions | Ionic strength: 0 Not defined / Label: 1 / pH: 5.5 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model: INOVA / Field strength: 500 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: molecular dynamics / Software ordinal: 1 / Details: used also for simulated annealing | ||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 200 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller












 PDBj
PDBj

 Amber
Amber