+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6c3g | ||||||
|---|---|---|---|---|---|---|---|
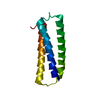
| Title | AMYLOID FORMING PEPTIDE KALGIS FROM TRANSTHYRETIN | ||||||
 Components Components | LYS-ALA-LEU-GLY-ILE-SER | ||||||
 Keywords Keywords | PROTEIN FIBRIL / amyloid / transthyretin / fibril | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.601 Å MOLECULAR REPLACEMENT / Resolution: 1.601 Å | ||||||
 Authors Authors | Saelices, L. / Sawaya, M.R. / Eisenberg, D.S. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2018 Journal: Protein Sci. / Year: 2018Title: Crystal structures of amyloidogenic segments of human transthyretin. Authors: Saelices, L. / Sievers, S.A. / Sawaya, M.R. / Eisenberg, D.S. #1:  Journal: J. Biol. Chem. / Year: 2015 Journal: J. Biol. Chem. / Year: 2015Title: Uncovering the Mechanism of Aggregation of Human Transthyretin. Authors: Saelices, L. / Johnson, L.M. / Liang, W.Y. / Sawaya, M.R. / Cascio, D. / Ruchala, P. / Whitelegge, J. / Jiang, L. / Riek, R. / Eisenberg, D.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6c3g.cif.gz 6c3g.cif.gz | 12.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6c3g.ent.gz pdb6c3g.ent.gz | 5.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6c3g.json.gz 6c3g.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  6c3g_validation.pdf.gz 6c3g_validation.pdf.gz | 396 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  6c3g_full_validation.pdf.gz 6c3g_full_validation.pdf.gz | 396 KB | Display | |
| Data in XML |  6c3g_validation.xml.gz 6c3g_validation.xml.gz | 2.9 KB | Display | |
| Data in CIF |  6c3g_validation.cif.gz 6c3g_validation.cif.gz | 3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/c3/6c3g https://data.pdbj.org/pub/pdb/validation_reports/c3/6c3g ftp://data.pdbj.org/pub/pdb/validation_reports/c3/6c3g ftp://data.pdbj.org/pub/pdb/validation_reports/c3/6c3g | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | x 10
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 588.718 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) Homo sapiens (human)#2: Chemical | ChemComp-SO4 / | #3: Chemical | ChemComp-GOL / | #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.53 Å3/Da / Density % sol: 19.71 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 3.5 Details: 0.1 M Citric acid pH 3.5, 2.0 M Ammonium sulfate, 25% Glycerol |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 24-ID-E / Wavelength: 0.9792 Å / Beamline: 24-ID-E / Wavelength: 0.9792 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Mar 1, 2013 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9792 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.6→12.566 Å / Num. obs: 871 / % possible obs: 95.6 % / Redundancy: 2.912 % / Biso Wilson estimate: 9.92 Å2 / CC1/2: 0.988 / Rmerge(I) obs: 0.138 / Rrim(I) all: 0.169 / Χ2: 1.154 / Net I/σ(I): 5.72 / Num. measured all: 2536 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.601→12.566 Å / SU ML: 0 / Cross valid method: THROUGHOUT / σ(F): 2 / Phase error: 24.35 MOLECULAR REPLACEMENT / Resolution: 1.601→12.566 Å / SU ML: 0 / Cross valid method: THROUGHOUT / σ(F): 2 / Phase error: 24.35
| ||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||
| Displacement parameters | Biso max: 32.44 Å2 / Biso mean: 14.45 Å2 / Biso min: 5.84 Å2 | ||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.601→12.566 Å
| ||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.6005→1.659 Å / Rfactor Rfree error: 0 / Total num. of bins used: 1
|
 Movie
Movie Controller
Controller















 PDBj
PDBj





