Entry Database : PDB / ID : 6a56Title AJLec from the Sea Anemone Anthopleura japonica AJLec Keywords / / / / Function / homology / / Biological species Anthopleura japonica (sea anemone)Method / / / Resolution : 1.2 Å Authors Unno, H. / Hatakeyama, T. Funding support Organization Grant number Country Japan Society for the Promotion of Science 15K06977
Journal : Sci Rep / Year : 2018Title : Identification, Characterization, and X-ray Crystallographic Analysis of a Novel Type of Lectin AJLec from the Sea Anemone Anthopleura japonica.Authors : Unno, H. / Nakamura, A. / Mori, S. / Goda, S. / Yamaguchi, K. / Hiemori, K. / Tateno, H. / Hatakeyama, T. History Deposition Jun 22, 2018 Deposition site / Processing site Revision 1.0 Jul 25, 2018 Provider / Type Revision 1.1 Aug 15, 2018 Group / Database references / Category / citation_authorItem _citation.country / _citation.journal_abbrev ... _citation.country / _citation.journal_abbrev / _citation.journal_id_CSD / _citation.journal_id_ISSN / _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.pdbx_database_id_DOI / _citation.pdbx_database_id_PubMed / _citation.title / _citation.year Revision 2.0 Jul 29, 2020 Group Atomic model / Data collection ... Atomic model / Data collection / Derived calculations / Non-polymer description / Polymer sequence / Structure summary Category atom_site / atom_site_anisotrop ... atom_site / atom_site_anisotrop / chem_comp / entity / entity_name_com / entity_poly / pdbx_branch_scheme / pdbx_chem_comp_identifier / pdbx_entity_branch / pdbx_entity_branch_descriptor / pdbx_entity_branch_link / pdbx_entity_branch_list / pdbx_entity_nonpoly / pdbx_molecule_features / pdbx_nonpoly_scheme / pdbx_struct_conn_angle / struct_asym / struct_conn / struct_site / struct_site_gen Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_asym_id / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.auth_seq_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.type_symbol / _atom_site_anisotrop.U[1][1] / _atom_site_anisotrop.U[1][2] / _atom_site_anisotrop.U[1][3] / _atom_site_anisotrop.U[2][2] / _atom_site_anisotrop.U[2][3] / _atom_site_anisotrop.U[3][3] / _atom_site_anisotrop.pdbx_auth_asym_id / _atom_site_anisotrop.pdbx_auth_atom_id / _atom_site_anisotrop.pdbx_auth_comp_id / _atom_site_anisotrop.pdbx_auth_seq_id / _atom_site_anisotrop.pdbx_label_asym_id / _atom_site_anisotrop.pdbx_label_atom_id / _atom_site_anisotrop.pdbx_label_comp_id / _atom_site_anisotrop.type_symbol / _chem_comp.formula / _chem_comp.formula_weight / _chem_comp.id / _chem_comp.mon_nstd_flag / _chem_comp.name / _chem_comp.pdbx_synonyms / _chem_comp.type / _entity.formula_weight / _entity.pdbx_description / _entity.type / _entity_poly.pdbx_seq_one_letter_code_can / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _struct_asym.entity_id Description / Provider / Type Revision 3.0 Nov 15, 2023 Group Atomic model / Data collection ... Atomic model / Data collection / Database references / Derived calculations / Refinement description / Structure summary Category atom_site / atom_site_anisotrop ... atom_site / atom_site_anisotrop / chem_comp / chem_comp_atom / chem_comp_bond / database_2 / pdbx_validate_peptide_omega / pdbx_validate_rmsd_angle / pdbx_validate_torsion / struct_conn / struct_ncs_dom_lim Item _atom_site.auth_atom_id / _atom_site.label_atom_id ... _atom_site.auth_atom_id / _atom_site.label_atom_id / _atom_site_anisotrop.pdbx_auth_atom_id / _atom_site_anisotrop.pdbx_label_atom_id / _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr2_label_atom_id / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_asym_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_asym_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Anthopleura japonica (sea anemone)
Anthopleura japonica (sea anemone) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  SAD / Resolution: 1.2 Å
SAD / Resolution: 1.2 Å  Authors
Authors Japan, 1items
Japan, 1items  Citation
Citation Journal: Sci Rep / Year: 2018
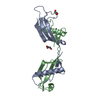
Journal: Sci Rep / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6a56.cif.gz
6a56.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6a56.ent.gz
pdb6a56.ent.gz PDB format
PDB format 6a56.json.gz
6a56.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads 6a56_validation.pdf.gz
6a56_validation.pdf.gz wwPDB validaton report
wwPDB validaton report 6a56_full_validation.pdf.gz
6a56_full_validation.pdf.gz 6a56_validation.xml.gz
6a56_validation.xml.gz 6a56_validation.cif.gz
6a56_validation.cif.gz https://data.pdbj.org/pub/pdb/validation_reports/a5/6a56
https://data.pdbj.org/pub/pdb/validation_reports/a5/6a56 ftp://data.pdbj.org/pub/pdb/validation_reports/a5/6a56
ftp://data.pdbj.org/pub/pdb/validation_reports/a5/6a56 Links
Links Assembly
Assembly
 Components
Components Anthopleura japonica (sea anemone) / References: UniProt: A0A2Z5WLM1*PLUS
Anthopleura japonica (sea anemone) / References: UniProt: A0A2Z5WLM1*PLUS X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Photon Factory
Photon Factory  / Beamline: BL-1A / Wavelength: 1.1 Å
/ Beamline: BL-1A / Wavelength: 1.1 Å Processing
Processing SAD / Resolution: 1.2→22.08 Å / Cor.coef. Fo:Fc: 0.976 / Cor.coef. Fo:Fc free: 0.968 / SU B: 1.42 / SU ML: 0.029 / Cross valid method: THROUGHOUT / ESU R: 0.037 / ESU R Free: 0.039 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
SAD / Resolution: 1.2→22.08 Å / Cor.coef. Fo:Fc: 0.976 / Cor.coef. Fo:Fc free: 0.968 / SU B: 1.42 / SU ML: 0.029 / Cross valid method: THROUGHOUT / ESU R: 0.037 / ESU R Free: 0.039 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller












 PDBj
PDBj








