Entry Database : PDB / ID : 5yspTitle Pyrophosphate-dependent kinase in the ribokinase family complexed with a pyrophosphate analog and myo-inositol TM0415 Keywords Function / homology Function Domain/homology Component
/ / / / / / / / / / Biological species Thermotoga maritima (bacteria)Method / / / Resolution : 1.7 Å Authors Nagata, R. / Fujihashi, M. / Miki, K. Funding support Organization Grant number Country Japan Society for the Promotion of Science 16J08482 Japan Society for the Promotion of Science 17H05439
Journal : Nat Commun / Year : 2018Title : Identification of a pyrophosphate-dependent kinase and its donor selectivity determinants.Authors : Nagata, R. / Fujihashi, M. / Sato, T. / Atomi, H. / Miki, K. History Deposition Nov 14, 2017 Deposition site / Processing site Revision 1.0 May 16, 2018 Provider / Type Revision 1.1 Nov 22, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / pdbx_struct_conn_angle / struct_conn Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr2_auth_seq_id
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 Thermotoga maritima (bacteria)
Thermotoga maritima (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å
MOLECULAR REPLACEMENT / Resolution: 1.7 Å  Authors
Authors Japan, 2items
Japan, 2items  Citation
Citation Journal: Nat Commun / Year: 2018
Journal: Nat Commun / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5ysp.cif.gz
5ysp.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5ysp.ent.gz
pdb5ysp.ent.gz PDB format
PDB format 5ysp.json.gz
5ysp.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads 5ysp_validation.pdf.gz
5ysp_validation.pdf.gz wwPDB validaton report
wwPDB validaton report 5ysp_full_validation.pdf.gz
5ysp_full_validation.pdf.gz 5ysp_validation.xml.gz
5ysp_validation.xml.gz 5ysp_validation.cif.gz
5ysp_validation.cif.gz https://data.pdbj.org/pub/pdb/validation_reports/ys/5ysp
https://data.pdbj.org/pub/pdb/validation_reports/ys/5ysp ftp://data.pdbj.org/pub/pdb/validation_reports/ys/5ysp
ftp://data.pdbj.org/pub/pdb/validation_reports/ys/5ysp

 Links
Links Assembly
Assembly
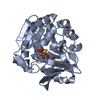

 Components
Components
 Thermotoga maritima (strain ATCC 43589 / MSB8 / DSM 3109 / JCM 10099) (bacteria)
Thermotoga maritima (strain ATCC 43589 / MSB8 / DSM 3109 / JCM 10099) (bacteria)

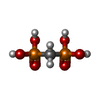







 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SPring-8
SPring-8  / Beamline: BL41XU / Wavelength: 1 Å
/ Beamline: BL41XU / Wavelength: 1 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj


