[English] 日本語
 Yorodumi
Yorodumi- PDB-5ur9: Enantiomer-Specific Binding of the Potent Antinociceptive Agent S... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5ur9 | ||||||
|---|---|---|---|---|---|---|---|
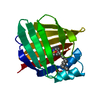
| Title | Enantiomer-Specific Binding of the Potent Antinociceptive Agent SBFI-26 to Anandamide transporters FABP5 | ||||||
 Components Components | Fatty acid-binding protein, epidermal | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN/INHIBITOR / inhibitor / LIPID BINDING PROTEIN-INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of prostaglandin biosynthetic process / regulation of retrograde trans-synaptic signaling by endocanabinoid / lipid transport across blood-brain barrier / retrograde trans-synaptic signaling by endocannabinoid / positive regulation of peroxisome proliferator activated receptor signaling pathway / negative regulation of D-glucose transmembrane transport / regulation of sensory perception of pain / phosphatidylcholine biosynthetic process / retinoic acid binding / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin ...regulation of prostaglandin biosynthetic process / regulation of retrograde trans-synaptic signaling by endocanabinoid / lipid transport across blood-brain barrier / retrograde trans-synaptic signaling by endocannabinoid / positive regulation of peroxisome proliferator activated receptor signaling pathway / negative regulation of D-glucose transmembrane transport / regulation of sensory perception of pain / phosphatidylcholine biosynthetic process / retinoic acid binding / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin / Signaling by Retinoic Acid / long-chain fatty acid transmembrane transporter activity / Triglyceride catabolism / epidermis development / long-chain fatty acid transport / fatty acid transport / postsynaptic cytosol / postsynaptic density, intracellular component / secretory granule membrane / fatty acid binding / lipid metabolic process / glucose metabolic process / azurophil granule lumen / glucose homeostasis / positive regulation of cold-induced thermogenesis / postsynaptic density / Neutrophil degranulation / synapse / lipid binding / glutamatergic synapse / extracellular exosome / extracellular region / nucleoplasm / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.19800343737 Å MOLECULAR REPLACEMENT / Resolution: 2.19800343737 Å | ||||||
 Authors Authors | Hsu, H.-C. / Li, H. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2017 Journal: Biochemistry / Year: 2017Title: The Antinociceptive Agent SBFI-26 Binds to Anandamide Transporters FABP5 and FABP7 at Two Different Sites. Authors: Hsu, H.C. / Tong, S. / Zhou, Y. / Elmes, M.W. / Yan, S. / Kaczocha, M. / Deutsch, D.G. / Rizzo, R.C. / Ojima, I. / Li, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5ur9.cif.gz 5ur9.cif.gz | 295.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5ur9.ent.gz pdb5ur9.ent.gz | 196 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5ur9.json.gz 5ur9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ur/5ur9 https://data.pdbj.org/pub/pdb/validation_reports/ur/5ur9 ftp://data.pdbj.org/pub/pdb/validation_reports/ur/5ur9 ftp://data.pdbj.org/pub/pdb/validation_reports/ur/5ur9 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5uraC  4lkpS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| 2 | 
| ||||||||||||
| 3 | 
| ||||||||||||
| 4 | 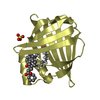
| ||||||||||||
| 5 | 
| ||||||||||||
| 6 | 
| ||||||||||||
| 7 | 
| ||||||||||||
| 8 | 
| ||||||||||||
| Unit cell |
| ||||||||||||
| Details | Monomer as determined by gel filtration |
- Components
Components
-Protein , 1 types, 8 molecules ABCDEFGH
| #1: Protein | Mass: 15467.732 Da / Num. of mol.: 8 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: FABP5 / Production host: Homo sapiens (human) / Gene: FABP5 / Production host:  |
|---|
-Non-polymers , 5 types, 373 molecules 








| #2: Chemical | ChemComp-8KS / ( #3: Chemical | ChemComp-SO4 / #4: Chemical | ChemComp-MYR / #5: Chemical | #6: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.87 Å3/Da / Density % sol: 57.2 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 0.1 M HEPES, pH 7.5, 2% polyethylene glycol 400, and 2.1 M ammonium sulfate 15 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X6A / Wavelength: 1 Å / Beamline: X6A / Wavelength: 1 Å |
| Detector | Type: ADSC QUANTUM 270 / Detector: CCD / Date: Mar 27, 2014 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.198→54.94 Å / Num. obs: 63855 / % possible obs: 92.5 % / Redundancy: 2.6 % / Biso Wilson estimate: 35.668891815 Å2 / CC1/2: 0.998 / Rmerge(I) obs: 0.056 / Net I/σ(I): 11.2 |
| Reflection shell | Resolution: 2.198→2.32 Å / Redundancy: 2.5 % / Rmerge(I) obs: 0.386 / Mean I/σ(I) obs: 2.3 / Num. unique obs: 8986 / CC1/2: 0.787 / % possible all: 89 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4LKP Resolution: 2.19800343737→40.7600279886 Å / SU ML: 0.290867111957 / Cross valid method: FREE R-VALUE / σ(F): 1.97336789233 / Phase error: 25.892558969
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 41.2183468876 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.19800343737→40.7600279886 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj










